Received: January 01, 1970
Published: January 01, 1970
Genet.Mol.Res. ():
Abbreviations
T1DM: Type 1 Diabetes Mellitus; T2DM: Type 2 Diabetes Mellitus; GES: Gestational Diabetes; MODY: Maturity Onset Diabetes of the Young.
Introduction
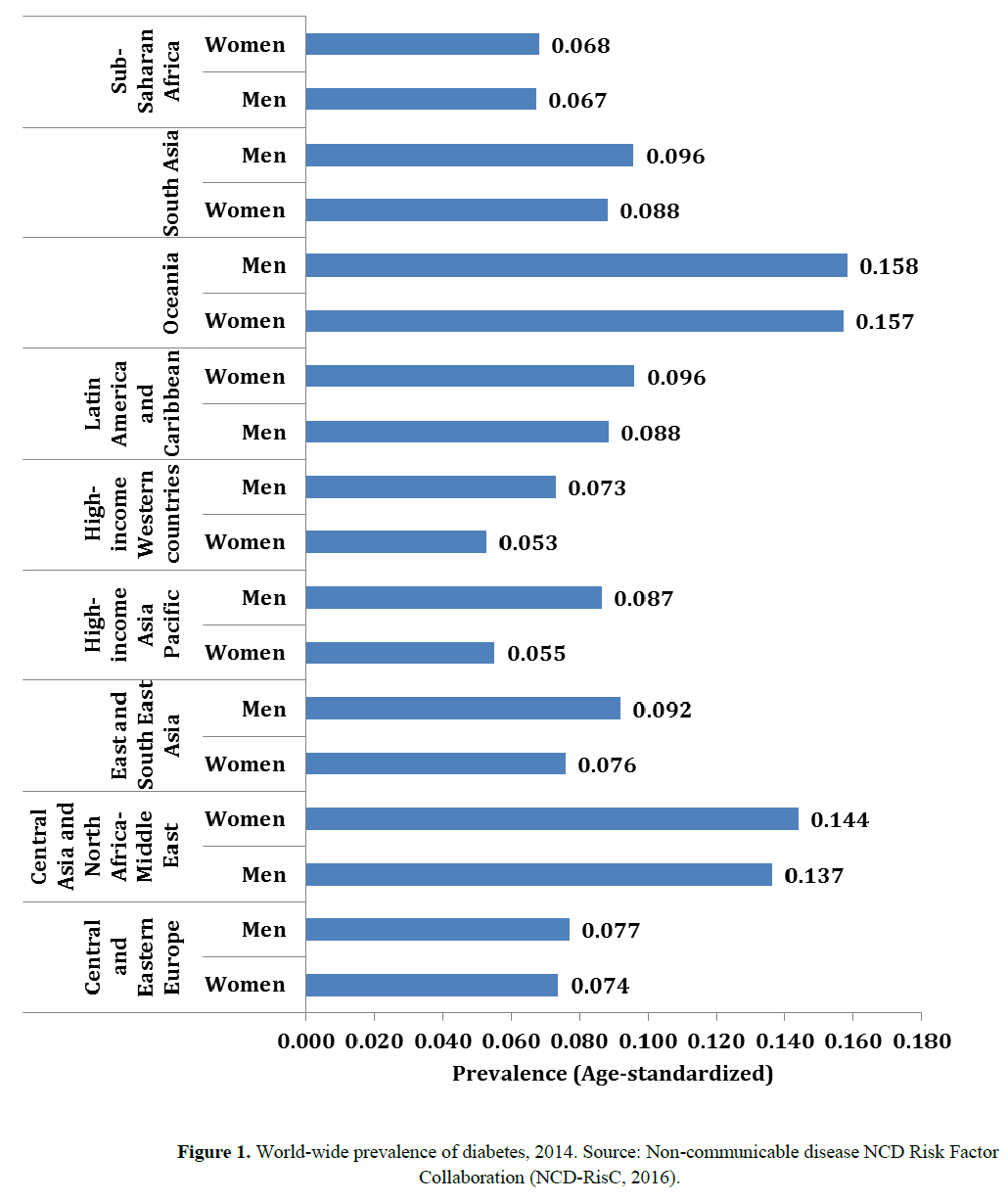
When fasting blood sugar level remains consistently above 126 mg/dL, then the condition is diagnosed as diabetes. Based on pooled analysis comprising of 751 population studies covering 146 countries representing 90% of worlds’ adult population, it was estimated that the age-standardized global prevalence of diabetes was 9% (among men) and 7.9% (among women) in 2014 (NCD-RisC, 2016). On a regional basis, the prevalence of diabetes was highest in both men and women of Oceania in 2014, crossing 15 percentage points. Among women, the lowest prevalence of 5.3% was observed in high-income western countries and among men; the lowest prevalence of age-standardized diabetes was 6.7% in sub-Saharan Africa (Figure 1). Co-existence of other clinical complications such as retinopathy, nephropathy, peripheral neuropathy, autonomic neuropathy, stroke and atherosclerosis leads to higher morbidity and mortality (American Diabetes Association, 2017; Forbes & Cooper, 2013; Mathers & Loncar, 2006; Shahi, Prasad, Imam, Abdul, & Jahangir, 2018).
Broadly, diabetes has been categorized as type-1 diabetes mellitus (T1DM), which is autoimmune mediated dysfunction of beta cells accounting for 5-10% of cases and type 2 diabetes mellitus (T2DM) which accounts for 90% of diabetes cases due to inadequate secretion of insulin (Atkinson, Eisenbarth, & Michels, 2014; Wild, Roglic, Green, Sicree, & King, 2004). The classification of diabetes has been revised based on latest developments in genetic understandings which revealed few monogenic disease forms including maturity onset diabetes of the young (MODY) and neonatal diabetes mellitus (NDM) accounting for 2-5% of cases around the globe. The GWAS reinforced our understanding of fundamental genetics by identifying new risk variants associated with various subtypes of diabetes. However, identifying the low frequency traits, emerging genetic risks and association with environmental factors remain challenging for the researchers. Particularly, the identification of locus and interpretation of functional roles of the gene variants is very challenging in case of multiple loci harboring common variants associated with T2DM (Cooper, Krawczak, Polychronakos, Tyler-Smith, & Kehrer-Sawatzki, 2013; Tallapragada, Bhaskar, & Chandak, 2015).
The present review summarizes the most recent outcomes from investigations of genetic mechanisms underlying the instigation, prognosis and manifestation of different subtypes of diabetes. The hypothesis remains to be tested as to whether any specific gene locus or group of gene loci exist that is/are common to different types of diabetes and if so, whether etiology of other forms of diabetes has any genetic association, either functionally or on positional basis, among the alleles associated with diabetes. It needs to be ascertained whether the manifestation of diabetes in general, is initiated by genetic polymorphism or alteration of metabolic pathway or both. Whether, there is any means of reversal of the process leading to permanent treatment or alleviation of the disease progression.
T1DM and the associated genetic determinants
T1DM is mostly due to the self-destruction of beta cells of pancreas; however, recent studies showed that the risk associated with at least one locus contributes to the familial clustering of T1DM. Advanced studies have found that clustering within major histocompatibility complex (MHC) located on chromosome 6 (6p21) is strongly (about 40%) associated with risk of T1DM. Recent genetic analyses and genome wide association studies (GWAS) revealed linkage of T1DM with many genes including insulin (INS), CTLA4, alongside HLA gene combinations (HLA-DQA1, HLA-DQB1and HLA-DRB1) (Pociot et al., 2010; Rich, 2006).
In type I diabetes mellitus (T1DM), the body self-immune system destroys the islets of Langerhans (beta cells) in the pancreas, mainly mediated by T-cell (Jerram & Leslie, 2017), resulting either in decreased level or no synthesis/secretion of insulin. Multiple genes are involved in the manifestation of T1DM, mostly auto-immune regulatory genes (Brownlie, Zamoyska, & Salmond, 2018). Single nucleotide polymorphisms (SNPs) in about 40 genetic loci mostly associated with insulin secretion were linked with T1DM (Barrett et al., 2009). Epigenetic variations caused by environmental and dietary factors also influence the disease prognosis mediated by autoimmune response. In the past few years, much of research was done on the genetic and inter-genic determinants of T1DM and their interactions with causal agents.
Abnormal environmental and dietary patterns result in the modification of the nutrient and nutrient derived metabolite profile. This affects the normal course of metabolic homeostasis leading to alterations in the content and proportion of chromatic remodeling metabolites. Consequently, the DNA methylation/demethylation as well as histone acetylation/deacetylation pattern alter, triggering aberrant gene expression, energy metabolism/balance and disruption of immune tolerance. The autoimmune response involving genes that are specific to insulin producing β-cells lead to instigation of T1DM (Wang, Long, Chang, Zhao, & Lu, 2018). Polygenic effect comprises of about 30 to 60 susceptible genes distributed over 50 regions was reported (Sanyoura, Philipson, & Naylor, 2018). Incidence of monogenic T1DM accounts for 1-2% of total cases. The genetic risk score of T1DM has the potential to identify monogenic autoimmunity in individuals. The specificity, precision and the probability of success in gene therapy for monogenic autoimmunity mediated diabetes is much higher than polygenic autoimmunity (Johnson et al., 2018). Recently, differentially methylated circulating insulin DNA in the serum derived from beta cells are now being considered to evaluate the prognosis of T1DM (Liu, Tan, & Liu, 2017).
Recent progress in genetic studies on T1DM
FOXP3 encodes transcription factor in T cell function (Hwang et al., 2018). Mutation in the gene leads to dysregulation of immunity ultimately resulting in early onset of T1DM. Impaired functioning of FOXP3 encoded regulatory T cells leads to generation of CD4+ and CD8+ T effector cells which cause beta cell destruction (Schallenberg & Kretschmer, 2018).
Interleukin 6 (IL-6) is a major pro-inflammatory cytokine in different types of tissues. Based on a systematic review and meta-data analysis of case control studies, it was revealed that serum content of IL-6 in T1DM patients was substantially higher than the control subject group and there was no correlation between the IL-6 level and age/ethnicity/disease duration (Chen et al., 2017). IL-6, apart from immunoregulatory effects also affects glucose homeostasis by causing insulin resistance in other cell types (Rehman et al., 2017).
A lot of variation exists in the etiology of T1DM. Insulin gene (INS) encodes pro-insulin which after post translational modification form insulin. Insulin is required for normal glucose homeostasis. Mutation in INS gene leads to misfolding resulting in hyperglycaemia and this phenomenon is commonly called as autoantibody-negative type 1 diabetes which is frequently observed in neonates and youth (Arunagiri et al., 2018; Barbetti et al., 2017).
ITPR3 gene encodes inositol 1,4,5-trisphosphate which is a receptor that mediates release of intracellular calcium. Polymorphism in this gene contributes to the risk of T1DM. In beta cells, when the voltage dependent Ca2+ channel opens the calcium ions enter into the cytoplasm thus enabling the secretion of insulin out of the cell. Since ITPR3 encodes receptor that regulates calcium transport, it is hypothesized that polymorphism in ITPR3 dysregulates insulin secretion (Rorsman & Ashcroft, 2018).
OAS1 gene encodes protein that activates RNAase that cleaves the viral RNA, thus conferring anti-viral property. The gene also affects other cellular process such as apoptosis, cell differentiation and gene regulation. The expression of the OAS1 gene was found to be significantly enhanced in T!DM patients when compared to normal subjects (Field et al., 2005) indicating that genetic response to viral infection may predispose the individual to T1DM. Single nucleotide polymorphism alters splicing (rs10774671) in OAS1 gene. Alternate splicing results in multiple transcript variants with different enzymatic activities. Substitution of serine/glycine in OAS1 exon 3 was identified as the most probable functional variant of the gene associated with increased enzymatic activity as well as with T1DM in case-sibling control study (Tessier et al., 2006). Later, it was observed that within OAS1 gene, A/G splice-site single nucleotide polymorphism (SNP; rs10774671) was associated with protective influence against T1DM among unaffected siblings (Smyth et al., 2006).
Protein tyrosine phosphatase N22 (PTPN22) gene encodes protein lymphoid tyrosine phosphatase (Lyp) that regulates signal transduction and play an important role in cellular differentiation. A missense mutation in the position 1858 from cytosine to thymidine in this gene leads to a change from Arg at position 620 to Trp which is associated with several autoimmune disorders that include T1DM. This allelic variation associated with autoimmunity was found to impair β-cell signaling which is characterized by proliferation deficit, reduced protein phosphorylation thus contributing to autoimmunity which affects β-cell maturation (Arechiga et al., 2009). Such intrinsic deficit in β-beta cell functioning could lead to T1DM.
Human leukocyte antigen (HLA) genes are the most impactful genes that encodes major histocompatibility complex (MHC) proteins involved in T1DM (Redondo, Steck, & Pugliese, 2018). HLA class II (and class I) genes have been identified as the most significant genetic risk factor associated with T1DM (Pociot et al., 2010). Polymorphism in non-HLA genes INS, PTPN22, CTLA4, and IL2RA are also genetic risk factors of T1DM. HLA-DQA1 and HLA-DQB1 genes transcribe and translate into two different proteins that bind together to form functional antigen-binding DQαβ heterodimer which displays foreign peptides to the immune system (Caillat-Zucman, 2017). The risk of auto-islet immunity mediated T1DM is increased by specific HLA DR/DQ allele. Highest risk for T1DM was found to be the DR34-DQ8 (DQ8 is DQA1*0301, DQB1*0302) heterozygous genotype (Aly et al., 2006). In another study, both HLA-DRB1*04 and -DRB1*03 frequencies were significantly higher among T1DM patients than in controls (Grubic et al., 2018). Variability in these genes is represented by several alleles that confer immunity to wide range of foreign antigens. Mutations in HLA genes override polygenic prevention of autoimmunity which makes human genome prone to T1DM (Noble & Valdes, 2011). The exact sequence of events and the mechanism of susceptibility are not clear as HLA refers to region in the genome constituted by several genes encoding different functional products. The reversal of the autoimmune process post onset is impractical. Therefore, prior diagnosis of the risk groups and prevention remains the effective strategy. Non DR/DQ loci within the MHC such as HLA-DP have been identified that have the potential to predict T1DM (Baschal & Eisenbarth, 2008).
Recent advances in T2DM genetics
Unlike T1DM, T2DM has no definitive cause; therefore, it is necessary to conduct multitude of studies to estimate the effects of risk factors such as age, sex, diet, physical activity/inactivity, familial history, genetics and ethnicity. The disease progression rate and associated complications are not similar across all T2DM patients; moreover, there are many factors involved in insulin resistance. This indicates that T2DM is a complex disease with coalition of metabolic disorders and combination of multiple intermediary traits. Hence, there is a strong rationale to change the pace of future studies to identify genetic association of T2DM (Ismail-Beigi et al., 2010; Thomas & Philipson, 2015; Zoungas et al., 2009). Genetic susceptibility of this multi-factorial disease has been studied as early as in 1980s which demonstrated a strong association of genetic components (Barnett, Eff, Leslie, & Pyke, 1981; Hansen & Pedersen, 2004). Studies conducted to identify candidate genes associated with T2DM have been employed and genes influencing the β cell function were identified (McCarthy, 2004). Genome-wide familial linkage-based approaches showed chromosomal positioning of genes such as HNF4A, CAPN10, ENPP1, ADIPOQ and TCF7L2, (Jose C Florez et al., 2012). GWAS focused on multiple common variants that are significant contributors in disease risk and progression. Subsequently, many consortiums were formed to perform large-scale studies to characterize the genetic basis of T2DM. DIAbetes Genetics Replication and Meta-analysis Consortium (DIAGRAM) is one of earliest study groups formed in the Europe which identified 31 novel loci that are associated with T2DM risk (http://diagram-consortium.org/about.html). Meta-Analyses of Glucose and Insulin-related traits Consortium (MAGIC) identified loci for fasting plasma glucose, fasting insulin, 2 h glucose, and HbA1c traits (Dupuis et al., 2010; Manning et al., 2012; Morris et al., 2012; Saxena et al., 2010; Scott et al., 2012; Voight et al., 2010) (https://www.magicinvestigators.org/). Genetic Investigation of Anthropometric Traits (GIANT) revealed hundreds of loci that are associated with obesity related/associated traits (Berndt et al., 2013; Locke et al., 2015; Randall et al., 2013; Shungin et al., 2015).
T2DM is a complex metabolic disorder involving multiple genes which are spatially distributed across different chromosomes and associated with diverse cellular signaling and metabolic pathways (Table 1). Environmental and dietary factors influence the risk of T2DM. However, genetic factors also play a significant role in the manifestation of T2DM. This is corroborated by the fact that a positive family history with diabetes confers a 2.5 fold greater risk for T2DM and about 15 to 25% of the first degree relatives of T2DM patients develop impaired glucose tolerance that may lead to T2DM (Stumvoll, Goldstein, & van Haeften, 2005). The risk of developing T2DM is about 38%, if one parent has T2DM (Pierce, Keen, & Bradley, 1995)and about 60% if both the parents are diabetic (Tattersal & Fajans, 1975). Description of some of the most relevant T2DM associated genes in terms of gene function under normal health condition, etiology, prevalence, risk alleles, type of genetic variation - their functional effects in disease condition are provided in this review.
| CHR . | GENE | DESCRIPTION | FUNCTION | LOCATION | START | END | LENGTH | EXONS | TYPE |
|---|---|---|---|---|---|---|---|---|---|
| 1 | PTPN22 | Protein Tyrosine Phosphatase, Non-Receptor Type 22 | Immunological | 1p13.2 | 113813811 | 113871761 | 57950 | 24 | T1DM |
| 2 | CTLA4 | Cytotoxic T-Lymphocyte Associated Protein 4 | Immunological | 2q33.2 | 203867788 | 203873960 | 6172 | 4 | T1DM |
| 2 | GCKR | Glucokinase Regulator | Metabolic | 2p23.3 | 27496839 | 27523689 | 26850 | 20 | MODY2 |
| 2 | IRS1 | Insulin Receptor Substrate 1 | Metabolic | 2q36.3 | 226731317 | 226798790 | 67473 | 2 | T1DM, GES |
| 2 | KLF11 | Kruppel Like Factor 11 | Transcription factor | 2p25.1 | 10043555 | 10054836 | 11281 | 6 | MODY7 |
| 2 | NEUROD1 | Neuronal Differentiation 1 | Transcription factor | 2q31.3 | 181668295 | 181680665 | 12370 | 4 | MODY6 |
| 3 | APPL1 | Adaptor Protein, Phosphotyrosine Interacting With PH Domain And Leucine Zipper 1 | Receptor | 3p14.3 | 57227737 | 57273471 | 45734 | 23 | MODY14 |
| 3 | CCR5 | C-C Motif Chemokine Receptor 5 (Gene/Pseudogene) | Immunological | 3p21.31 | 46370142 | 46376206 | 6064 | 3 | T1DM |
| 3 | IGF2BP2 | Insulin Like Growth Factor 2 Mrna Binding Protein 2 | Genetic | 3q27.2 | 185643131 | 185825056 | 181925 | 24 | GES |
| 3 | PPARG | Peroxisome Proliferator Activated Receptor Gamma | Genetic | 3p25.2 | 12287485 | 12471054 | 183569 | 14 | T2DM |
| 4 | WFS1 | Wolframin ER Transmembrane Glycoprotein | Metabolic | 4p16.1 | 6260368 | 6303265 | 42897 | 10 | T2DM |
| 6 | CDKAL1 | CDK5 Regulatory Subunit Associated Protein 1 Like 1 | Genetic | 6p22.3 | 20534457 | 21232404 | 697947 | 22 | GES |
| 6 | HLA-DQA1 | Major Histocompatibility Complex, Class II, DQ Alpha 1 | Immunological | 6p21.32 | 32637406 | 32654846 | 17440 | 6 | T1DM |
| 6 | HLA-DQB1 | Major Histocompatibility Complex, Class II, DQ Beta 1 | Immunological | 6p21.32 | 32659464 | 32666689 | 7225 | 6 | T1DM |
| 6 | HLA-DRB1 | Major Histocompatibility Complex, Class II, DR Beta 1 | Immunological | 6p21.32 | 32578769 | 32589836 | 11067 | 6 | T1DM |
| 6 | ITPR3 | Inositol 1,4,5-Trisphosphate Receptor Type 3 | Metabolic | 6p21.31 | 33621333 | 33696574 | 75241 | 60 | T1DM |
| 6 | SUMO4 | Small Ubiquitin-Like Modifier 4 | Metabolic | 6q25.1 | 149400359 | 149401046 | 687 | 1 | T1DM |
| 7 | GCK | Glucokinase | Metabolic | 7p13 | 44143213 | 44189439 | 46226 | 15 | GES |
| 7 | GCK | Glucokinase | Enzyme | 7p13 | 44143213 | 44189439 | 46226 | 15 | MODY2 |
| 7 | IL6 | Interleukin 6 | Immunological | 7p15.3 | 22725889 | 22732002 | 6113 | 6 | T1DM |
| 7 | PAX4 | Paired Box 4 | Transcription factor | 7q32.1 | 127610292 | 127618192 | 7900 | 12 | MODY9 |
| 8 | BLK | BLK Proto-Oncogene, Src Family Tyrosine Kinase | Receptor | 8p23.1 | 11493991 | 11564604 | 70613 | 15 | MODY11 |
| 9 | CEL | Carboxyl Ester Lipase | Enzyme | 9q34.13 | 133061978 | 133071863 | 9885 | 11 | MODY8 |
| 10 | IL2RA | Interleukin 2 Receptor Subunit Alpha | Immunological | 10p15.1 | 6010694 | 6062370 | 51676 | 8 | T1DM |
| 10 | TCF7L2 | Transcription Factor 7 Like 2 | Genetic | 10q25.2-q25.3 | 112950220 | 113167678 | 217458 | 20 | GES |
| 11 | INS | Insulin | Metabolic | 11p15.5 | 2159779 | 2161209 | 1430 | 3 | T1DM |
| 11 | INS | Insulin | Hormone | 11p15.5 | 2159779 | 2161209 | 1430 | 3 | MODY10 |
| 11 | KCNJ11 | Potassium Voltage-Gated Channel Subfamily J Member 11 | Metabolic | 11p15.1 | 17385246 | 17389331 | 4085 | 3 | GES |
| 11 | KCNJ11 | Potassium Voltage-Gated Channel Subfamily J Member 11 | Transporter | 11p15.1 | 17385246 | 17389331 | 4085 | 3 | MODY12 |
| 11 | KCNQ1 | Potassium Voltage-Gated Channel Subfamily Q Member 1 | Metabolic | 11p15.5-p15.4 | 2444991 | 2849110 | 404119 | 17 | GES |
| 11 | MTNR1B | Melatonin Receptor 1B | Metabolic | 11q14.3 | 92969623 | 92986241 | 16618 | 4 | GES |
| 12 | HNF1A | HNF1 Homeobox A | Genetic | 12q24.31 | 120977683 | 121002512 | 24829 | 9 | T1DM, T2DM, MODY3 |
| 12 | OAS1 | 2'-5'-Oligoadenylate Synthetase 1 | Immunological | 12q24.13 | 112906777 | 112932190 | 25413 | 8 | T1DM |
| 13 | IRS2 | Insulin Receptor Substrate 2 | Metabolic | 13q34 | 109753837 | 109786567 | 32730 | 2 | T2DM |
| 13 | PDX1 | Pancreatic And Duodenal Homeobox 1 | Transcription factor | 13q12.2 | 27919982 | 27926314 | 6332 | 2 | MODY4 |
| 17 | HNF1B | HNF1 Homeobox B | Transcription factor | 17q12 | 37686431 | 37745078 | 58647 | 11 | MODY5 |
| 20 | HNF4A | Hepatocyte Nuclear Factor 4 Alpha | Genetic | 20q13.12 | 44355801 | 44432845 | 77044 | 13 | T2DM, GES |
| 20 | HNF4A | Hepatocyte Nuclear Factor 4 Alpha | Transcription factor | 20q13.12 | 44355801 | 44432845 | 77044 | 13 | MODY1 |
| X | FOXP3 | Forkhead Box P3 | Immunological | Xp11.23 | 49250436 | 49264932 | 14496 | 12 | T1DM |
Table 1. Genetic features of different types of diabetes.
Peroxisome proliferator-activated receptor gamma (PPAR-γ) gene encodes nuclear receptor protein that regulates adipocyte differentiation and glucose homeostasis. PPAR gamma was found to be down regulated in T2DM and in colon cancer as well (Lecarpentier, Claes, Vallée, & Hébert, 2017). PPAR gamma is activated by certain fatty acids, prostanoids, and thiazolidinediones (Olefsky, 2000). After activation by a ligand, the nuclear receptor protein regulates the transcription of other genes associated with fat partitioning into adipocyte and free fatty acid oxidation thus lowering free fatty acid content in the blood. Polymorphic variant of the Pro12Ala in PPAR gamma is highly prevalent (75%) in white people (Stumvoll et al., 2005). Heterozygous negative dominant mutations causing functional loss in PPAR-γ leads to dyslipidemia, insulin resistance and hyperglycaemia (Agostini et al., 2018). Point mutations in PPAR-γ lead to a gradation of metabolic response. Supplementation of flaxseed oil for about three months was found to significantly improve PPAR-γ gene expression in diabetic patients with coronary heart disease (Hashemzadeh et al., 2017). PPAR-γ protein has the potential to serve as anti-diabetic molecular target (Kerru, Singh-Pillay, Awolade, & Singh, 2018).
IRS1 gene encodes protein that acts as a substrate for insulin receptor tyrosine kinase. It bears an important role in insulin stimulated signal transduction pathway (Copps & White, 2012). A gene expression meta-analysis has revealed that IRS1 is a tissue non-specific gene which is significantly associated with insulin response. Genetic epidemiology of T2DM in Mexico revealed that IRS1 gene polymorphism was significantly related to the increased risk of T2DM in which SNP in at least two out of total five loci were significant (García-Chapa, Leal-Ugarte, Peralta-Leal, Durán-González, & Meza-Espinoza, 2017). Gly971Arg substitution in this gene leads to impaired signaling and thus insulin resistance. Recently, it was observed that up-regulated hepatic mRNA (miR-222) in the insulin resistant state represses IRS1 gene expression leading to impaired insulin signaling (Ono et al., 2018).
IRS2 gene encodes protein that functions as an adaptor to insulin receptor in order to activate kinase cascade in insulin signaling. Therefore, repression of IRS2 gene leads to insulin resistance (Zhang & Sun, 2009). Recently, it was observed that hyperglycaemia alters expression of IRS proteins in nerve cells leading to impaired insulin signal cascade and diabetic peripheral neuropathy (Manu, Rachana, & Advirao, 2017). Among the recent findings, polymorphism in Gly1057Asp of IRS-2 gene was found to be associated with T2DM (Ay et al., 2018).
Recently, Kleiner et al. have reported that loss of function mutation in SLC30A8 which encodes protein that transports zinc into insulin containing secretory granules in β-cells may exert beneficial influence on insulin secretion under hyperglycaemic conditions (Kleiner et al., 2018). Zhong et al. have observed that silencing of DDX1 gene (encoding multifunction protein) impairs calcium influx and insulin secretion in β-cells (Zhong, Li, Zhou, Xu, & Wang, 2018). Juan-Mateu et al. have noted that SRp55 (a splicing factor) regulated by GLIS3 functions for splicing of genes involved in cell survival, insulin secretion, and c-Jun N-terminal kinase (JNK) signaling. Depletion of SRp55 inhibits β-cell mitochondrial functioning, decreases insulin secretion and triggers β-cell apoptosis (Juan-Mateu et al., 2018).
WFS1 gene encodes wolframin glycoprotein located in endoplasmic reticulum membrane which regulates the amount of calcium within the cell thus enabling endoplasmic reticulum to function in protein folding. Allelic mutation in WFS1 gene causes autosomal recessive wolfram syndrome characterized by non-auto-immune diabetes (Bansal, Boehm, & Darvasi, 2018). Hofmann et al. have emphasized that wolfram syndrome is caused by reduced level of protein expression rather than mutant wolframin because of rapid decay of WFS1 nonsense transcripts as well as reduced content and/or reduced half-life of steady state wolframin in case of missense mutation R629W (Hofmann, Philbrook, Gerbitz, & Bauer, 2003). Bansal et al. identified a low frequency missense coding variant (c.1672C>T, p.R558C) in the WFS1 gene among Jewish individuals that causes wolfram syndrome (Bansal et al., 2018). Li et al. also identified a mutation in WFS1 gene in exon 8 (c.1760G > A) that caused diabetes in a Chinese patient (Li et al., 2018). Similarly, several mutations have been identified by Rigoli et al. that were linked to T2DM (Rigoli, Bramanti, Di Bella, & De Luca, 2018). Frank et al. have established a strong association between SNP in WFS1 locus, rs10010131, with risk of T2DM among European population (Franks et al., 2008). Similarly, Minton et al. have also observed that WFS1 gene variant influences susceptibility to T2DM among UK population and R456-H611 haplotype was highly frequent in type 2 diabetic subjects (Minton et al., 2002).
HNF1A encodes transcription factor that expresses pancreatic and hepatic genes. Mutation in HNF1A accounts for upto 3% of diabetic cases among children (Rama Chandran et al., 2018). The genetic variant of HNF1A gene is among the 40 confirmed variant loci associated with T2DM (McCarthy, 2010). Nuli et al. reported that HNF1A gene variants of rs2464196, rs1169288 and rs1169289 might interact with smoking and dietary factors in conferring risk of T2DM in the Uyghur population (Nuli et al., 2018). The variant rs2259816 (HNF1A) was found to be significantly associated with risk of coronary heart disease among T2DM patients (De Rosa et al., 2018). However, some studies have noted that the genetic variants of HNF1A affecting insulin secretion were found to be only associated with T2DM in non-obese individuals. Based on whole-exome sequencing, a low frequency missense variant of HNF1A was found to be associated with T2DM prevalence in population of Latinos (Estrada et al., 2014). Meta-analytical studies revealed two missense SNPs of A98V and I27L associated with T2DM that were reported to manifest a decrease in insulin secretion (Holmkvist et al., 2006).
HNF4A encodes protein that regulates the transcription of HNF1A thus regulating upstream hepatic gene expression. Although the variants of HNF4A are rare (Huerta-Saenz, Saunders, & Yan, 2018), they were identified as putative causal factor for T2DM affecting β-cell functioning (Kooner et al., 2011; Sladek et al., 2007). Allelic variant of rs4812829 in HNF4A was strongly associated with β-cell functioning and proinsulin, whereas rs1800961 was linked with lipid and proinsulin suggesting insulin secretion and insulin resistance mechanisms (Udler et al., 2018). The risk allele of rs4812829 in HNF4A was identified as an SNP that is strongly associated with T2DM among South Asian population (Kooner et al., 2011). However, it was observed that the variants of HNF4A were associated with a very small percentage of T2DM risk (J C Florez, 2008).
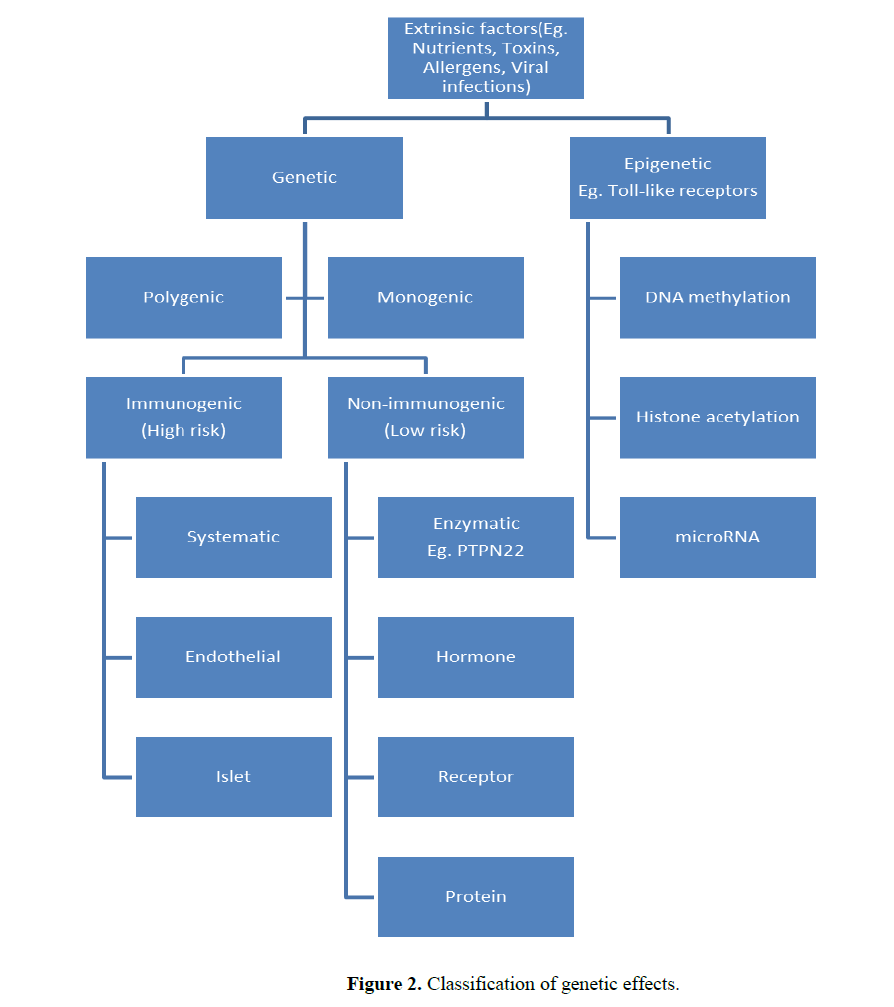
The genetic similarity between T1DM and T2DM is that both belong to polygenic variations and are caused due to the interactions between genetic and environmental factors. The basic determinants of diabetic condition and their effects are presented in Figure 2.
Genetics challenges of monogenic diabetes
The monogenic diabetes is a rare heterogeneous group of disorders arising due to defects in single gene, where hyperglycemia is observed due to defects in insulin secretion or decrease in beta cell mass or both (Schwitzgebel, 2014; Thomas & Philipson, 2015; Wiley, 2016). MODY is a form of monogenic diabetes, which is diagnosed at the childhood or early adulthood (before 25 years of age) stages. MODY is clinically and genetically heterogeneous, non-insulin dependent and familial diabetes which is caused due to the autosomal dominance inheritance. According to American Diabetes Association (ADA) guidelines, monogenic diabetes can be considered, when an individual does not have T1DM and T2DM features and diagnosed before 6 months of life with a strong familial history of diabetes, and stable/mild elevated fasting blood glucose levels (American Diabetes Association, 2017). Around 13 genes are associated with risk of MODY and mutations in glucose sensor enzyme glucokinase and 5 transcription factors are involved in regulation of adult β-cell function and a role in MODY progression. It is estimated that the mutations in GCK and HNF1 homeobox A (HNF1A) accounts for approximately 50% cases and mutations in HNF4A and HNF1B accounts for 10% cases while the genetic causes for few sub-types of MODY remains unknown (Ellard, Bellanne-Chantelot, & Hattersley, 2008; Rubio-Cabezas et al., 2014). Recent studies employed high-throughput sequencing technologies sequenced the genes associated with diabetes and obesity on large samples to identify target gene candidates (Bonnefond et al., 2014; Froguel, 2015; Molven & Njolstad, 2011). A Illumina high-throughput sequencing technology study sequenced a large set of genes to identify the association between monogenic diabetes and T2DM. In this study, on an overall basis, 22 genes were found to be associated with monogenic forms of diabetes which includes 13 MODY genes, 6 genes associated with recessive diseases that include diabetes as a phenotype and 3 genes within which heterozygous mutations lead to diabetes (Bonnefond et al., 2014; Ellard et al., 2013; Flannick et al., 2013). Next generation sequencing, whole genome sequencing and exome sequencing provided more insights into low frequency risk variants and disease causing mechanisms (Claussnitzer et al., 2014; Flannick et al., 2014; Majithia et al., 2014; Steinthorsdottir et al., 2014).
Monogenic diabetes forms are rare and caused due to mutation in single gene and in many cases the mutations are inherited from one or both parents. MODY and NDM are two common subtypes of monogenic diabetes (Barbetti & D’Annunzio, 2018). Most recent GWA studies showed that there are loci and mechanisms that are common in MODY and T2DM (Biesecker, 2018). Mapping function approaches found that cis-regulation is a common mechanism associated with affected transcriptional enhancers and silencers (Fogarty, Cannon, Vadlamudi, Gaulton, & Mohlke, 2014; Fogarty, Panhuis, Vadlamudi, Buchkovich, & Mohlke, 2013; López Rodríguez et al., 2017). There are different genes associated with MODY and few of them have a strong correlation with other forms of diabetes. In addition, clinical features of MODY depend on the type of gene mutation (Gardner & Tai, 2012). It is believed that in certain types of mutations, there are no symptoms of diabetes and or there is no development of associated complications (https://www.who.int/genomics/about/Diabetis-fin.pdf). As of now, about 14 genes responsible for the onset of MODY were reported and few of them also pose a risk for NDM (Table 1).
According to Biesecker, T2DM or adult onset diabetes is associated with several genes with common susceptibility variants which can lead to single gene or Mendelian form of diabetes, for example glucokinase (GCK) related T2DM (Biesecker, 2018). Therefore, such rare forms of T2DM need to be differentiated from single gene MODY. The diagnosis of MODY which is an atypical autoimmune form of diabetes is challenging due to clinical and genetic heterogeneity. Recently, Pace et al. reported a rare missense mutation in exon 8 of HNF1β (p.Arg527Gln) in obese Maltese women with autoimmune diabetes revealing monogenic diabetes could contribute substantially to disease burden (Pace, Craus, Felice, & Vassallo, 2018).
HNF1β gene mutation leads to MODY5 which is a multi-systemic syndrome and a monogenic form of diabetes. In adults, HNF1A-MODY is considered as the most frequently observed form of monogenic diabetes which can be potentially misdiagnosed as T1DM or T2DM; hence, genetic diagnosis is required. Pavic et al. attempted to estimate the prevalence of HNF1A-MODY among adults below 45 years of age with HNF1A allele variants in Croatia (Pavic et al., 2018). The study revealed the prevalence rate of HNF1A-MODY was 66 cases per million which was similar to other European countries.
Pollak et al. have recently reported that GCK gene sequencing resulted in accurate molecular diagnosis of MODY2 and facilitated adequate medical treatment in an affected Chilean family (Pollak C et al., 2017). Khelifa et al. have reported that GCK and HNF1A mutations are most frequently observed among Caucasians and attempted to evaluate the etiology of MODY in Tunisia (Ben Khelifa et al., 2018). After screening for mutations in GCK, HNF1A, HNF4A and INS genes, it was found that there were no mutations in HNF1A and INS genes suggesting the probable involvement of other unidentified genes (Ben Khelifa et al., 2018).
Genetic variants of gestational diabetes
Gestational diabetes is a condition where normo-glycemic women prior to pregnancy develop glucose intolerance in late gestational period resulting in subclinical manifestation of metabolic dysfunction. About 50 to 60% decrease in the insulin sensitivity can be observed during normal pregnancy (Catalano, 2014). Decreased β-cell functioning (lower secretion of insulin) and the lack of insulin sensitivity or increased insulin resistance in peripheral tissues leads to hyperglycemia in such women. In normal women with glycemic tolerance, this is a transient phase and gets reversed after giving birth. However, in women with gestational diabetes, the inflammatory response and low insulin sensitivity continues post-partum due to reduced tyrosine phosphorylation of insulin receptor substrate 1 (IRS1). Gene array studies have emphasized that alteration of gene expression pattern in lipid metabolism leading to hyperlipidemia were mainly involved in development of gestational diabetes (Catalano, 2014). Gestational diabetes manifests in the late second trimester period. The prevalence of gestational diabetes is about 1 to 3% of all the pregnancies. Women with previous history of gestational diabetes have a high risk of developing non-insulin dependent diabetes mellitus (NIDDM) (Damm, 1998). In a recent study comprising of 262 Euro-Brazilian pregnant women, it was observed that polymorphism of rs17567 (Gln279Arg) in Metalloproteinase 9 Gene (MMP9) was associated with gestational diabetes (Costa et al., 2018). Recently, Tagoma et al. have found that about 15 miRNA involved in 41 pathways were up-regulated in women with gestational diabetes (Tagoma, Alnek, Kirss, Uibo, & Haller-Kikkatalo, 2018). Among them, pathways associated with fatty acid biosynthesis and metabolism were the most relevant to gestational diabetes. The study concludes that increased plasma miR-195-5p miRNA is the cause of gestational diabetes development. In addition to these, Yan et al. have demonstrated that up-regulation and down regulation of circular mRNA in placental villi of gestational diabetes patients, most of which had small miRNA binding sites, could have potential role in development of gestational diabetes (Yan et al., 2018).
The gene GCKR (glucokinase regulator) encodes a 68kD sugar isomerase (SIS) regulatory protein that inhibits glucokinase activity in hepatic and pancreatic β-cells by binding with the enzyme and relocating it to nucleus. The chromosomal location and gene properties are presented in Table 1. The role of GCKR has been strongly associated with T2DM. Based on a case control meta-analytical study including genotype data of 1122 women, Jamalpour et al. observed a strong association of GCKR rs780094-C with higher risk of gestational diabetes (Jamalpour, Zain, Mosavat, Mohamed, & Omar, 2018).
Chimerin has been regarded as one of the metabolic mediator in the development of insulin resistance. Hasanvand et al. observed an association between Chemerin SNPs rs4721 and the risk for developing gestational diabetes among Iranian women (Hasanvand et al., 2018). TCF7L2 (transcription factor 7 like 2) encodes for a transcription factor involved in Wnt signaling pathway regulating blood glucose homeostatis. Franzago et al. observed that genetic variant - rs7903146 (C > T) in TCF7L2 gene was strongly associated with the risk of developing gestational diabetes while polymorphism in PPARG2 gene was found to be strongly correlated with lipid parameters (Franzago et al., 2018).
Macrophage migration inhibitory factor (MIF) gene encodes lymphokine involved in macrophage function and inflammation. Zheng et al. have revealed that enhanced expression of MIF gene in placental tissue correlated with insulin resistance and incidence of gestational diabetes among Chinese Han population (Zheng et al., 2017). Alpha-2-adrenergic receptors (ADRA2A) gene encodes receptors that regulate neurotransmitter release from nerves. Kawai et al. have revealed that genetic variation in ADRA2A gene is independently associated with development of risk of gestational diabetes among Caucasian women (Kawai et al., 2017).
Kang et al. measured DNA methylation at 841,573 CpG sites and identified 200 loci of the genes in maternal and cord blood which were methylated differently in pregnant women with gestational diabetes, among which inflammatory pathways were identified (Kang, Lee, Li, Hsu, & Lin, 2017). Solute carrier family 6 member 4 (SLC6A4) encodes membrane protein that transports serotonin neurotransmitter from synaptic spaces into presynaptic neurons for reutilization. Blazevic et al. have noticed that DNA methylation of the fetal SLC6A4 gene is related to metabolic state in pregnant women. DNA methylation across seven loci was negatively correlated with plasma glucose levels at 28th week of gestation. Paired box gene (PAX) encodes a family of transcription factors which are involved in differentiation and development of pancreatic β-ells. Xu et al. have observed that polymorphism in rs10229583 near paired box gene (PAX) is associated with gestational diabetes among Chinese women wherein GG genotype frequency was significantly different between gestational diabetes and normal subjects (Xu et al., 2018). Based on a study comprising of 204 pregnant women with gestational diabetes, Tarnowski et al. have found that polymorphisms in pro-inflammatory cytokine genes (IL18 rs187238 and rs1946518) were associated with higher risk of gestational diabetes (Tarnowski et al., 2017).
Genome wide association studies of diabetes condition
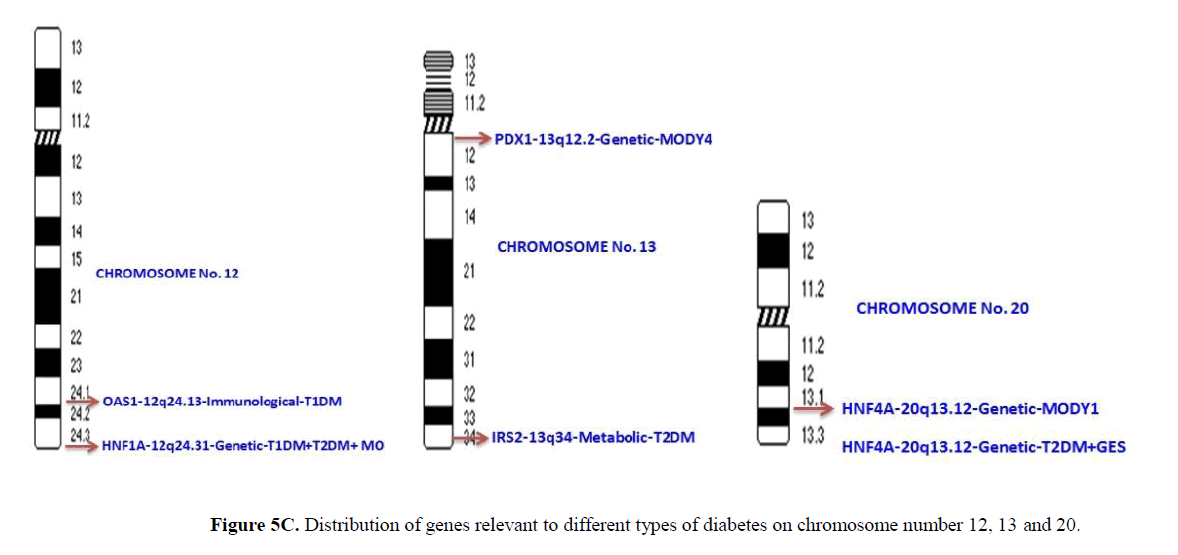
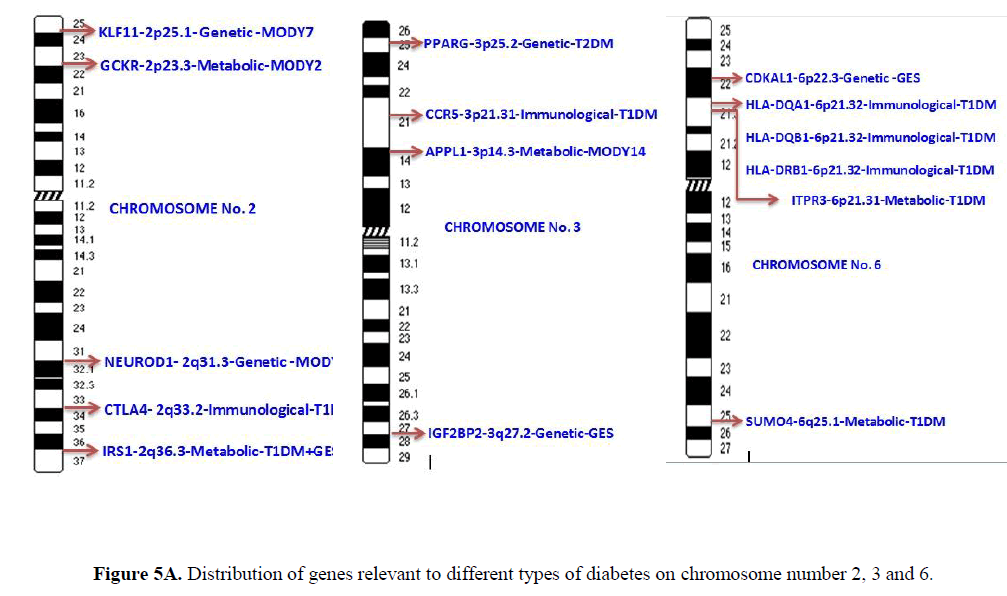
Genomic studies have revealed that mutations in genes distributed across chromosomes were associated with different types of diabetes such as T1DM, T2DM, and gestational diabetes and MODY (Kota, Meher, Jammula, Kota, & Modi, 2012). The distribution of the genes involved in development of different types of diabetes is presented in Table 1. Chromosome number 6 and 11 harbors the maximum number of genes (6), each associated with different types of diabetes. This is followed by chromosome number 2 carrying 5 diabetes related genes. Chromosome 3 has 4 genes while chromosome 10, 12, 13 and 20 have 2 genes each. Chromosome 1, 4, 8, 9, 17 and X chromosome contain 1 gene each.
Out of all the 39 genes evaluated, 10 genes were related to immunological function, 17 genes were involved in metabolic function and 12 genes have genetic regulatory roles. The length of the gene ranged from a minimum of 687 base pairs for small ubiquitin-like modifier 4 (SUMO4) gene located on the long arm of chromosome number 6 to maximum of 697947 base pairs for CDK5 regulatory subunit associated protein 1 like 1 (CDKAL1) gene located on small arm of chromosome number 6. The number of exons ranged from a minimum of 1 exon in SUMO4 to a maximum of 60 exons in ITPR3 gene located on small arm of chromosome number 6. Except chromosome number 1 and X chromosome, all other chromosomes had genes related to more than one type of diabetes. A total of 15 genes were linked to T1DM while only 5 genes were associated with T2DM. Gestational diabetes was related to 9 genes while 14 genes were related to MODY. It is interesting to note that mutation in IRS1 gene located on chromosome number 2 was responsible for both T1DM and gestational diabetes. HNF1A gene on long arm of chromosome number 12 was linked to three different types of diabetes namely T1DM, T2DM and MODY3. HNF4A gene located on the long arm of chromosome number 20 was linked to two different types of diabetes namely T2DM and gestational diabetes. Maximum heterogeneity with regard to the genes related to different types of diabetes was found with chromosome number 3 that contained genes related to all four types of diabetes (Table 1).
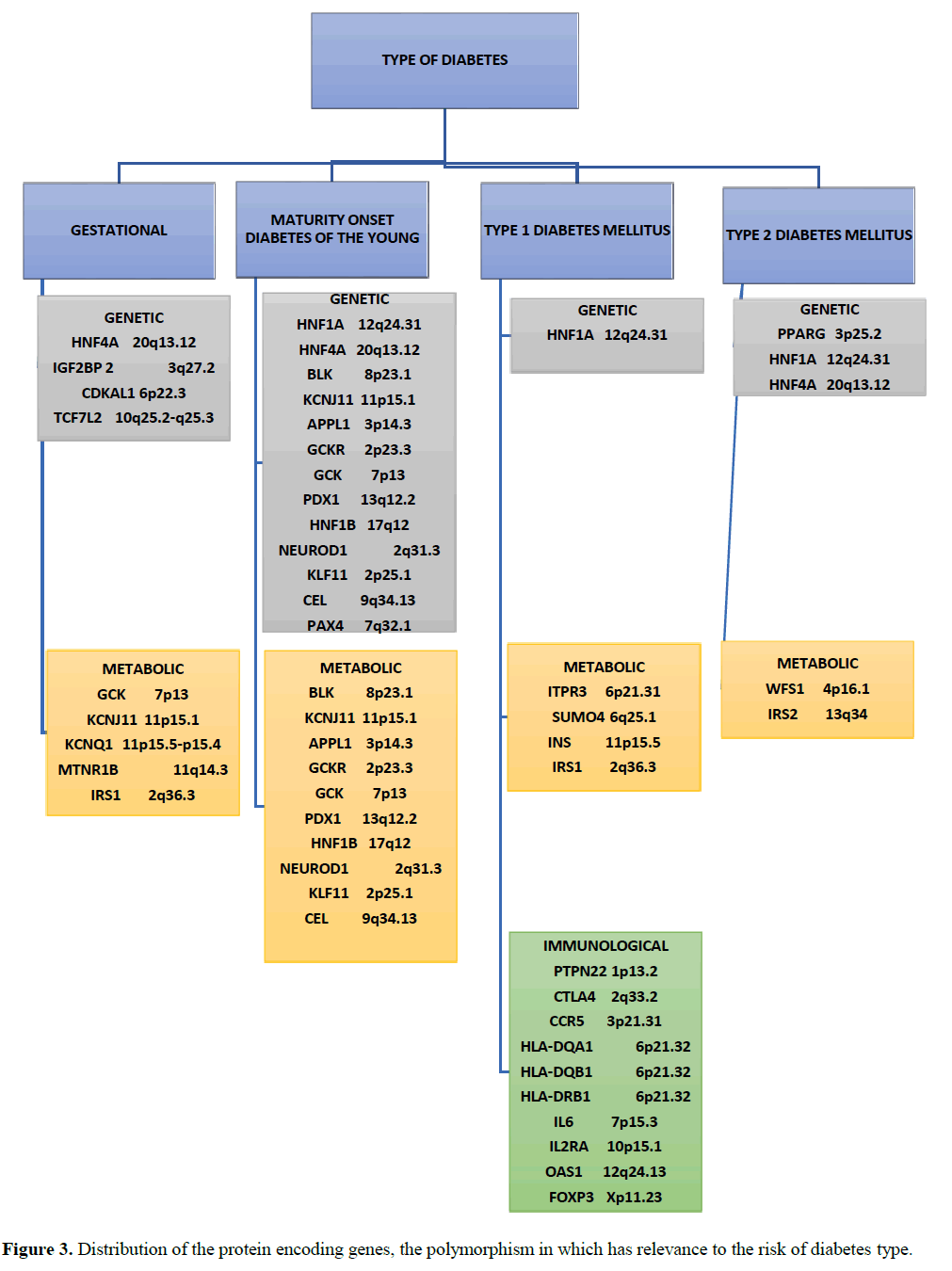
Considering the genes encoding protein with genetic role in the manifestation of the risk in different types of diabetes, it was found that a greater number of genes were associated with MODY followed by gestational diabetes, T2DM and T1DM (Figure 3). Single nucleotide polymorphism in about thirteen genes were relevant for MODY while the number of genes relevant to the manifestation of gestational diabetes, T2DM and T1DM were 4, 3 and 1, respectively. It is interesting to note that HNF4A gene located in the position 20q13.12 was common for gestational diabetes, MODY and T2DM.
HNF1A gene located in the position of 12q24.31 was common for the risk of MODY, T1DM and T2DM. Considering the genes encoding protein that have metabolic function, it was observed that a maximum of 10 genes were associated with the risk of MODY followed by 5 genes related to the risk of gestational diabetes, 4 genes relevant to the risk of T1DM and 2 genes related to T2DM. GCK gene located on 7p13 position, KCNJ11 located at 11p15.1 was common for gestational diabetes and MODY. IRS1 located at 2q36.3 was common for gestational diabetes and T2DM. With regard to the genes encoding protein having immunological function, a maximum of 10 genes were associated with the risk of T1DM. It is interesting to note that among major protein encoding genes with immunological role in which SNP has profound influence on the risk of diabetes were all solely related to T1DM. Probably, this is the reason why T1DM is also termed as autoimmune disease.
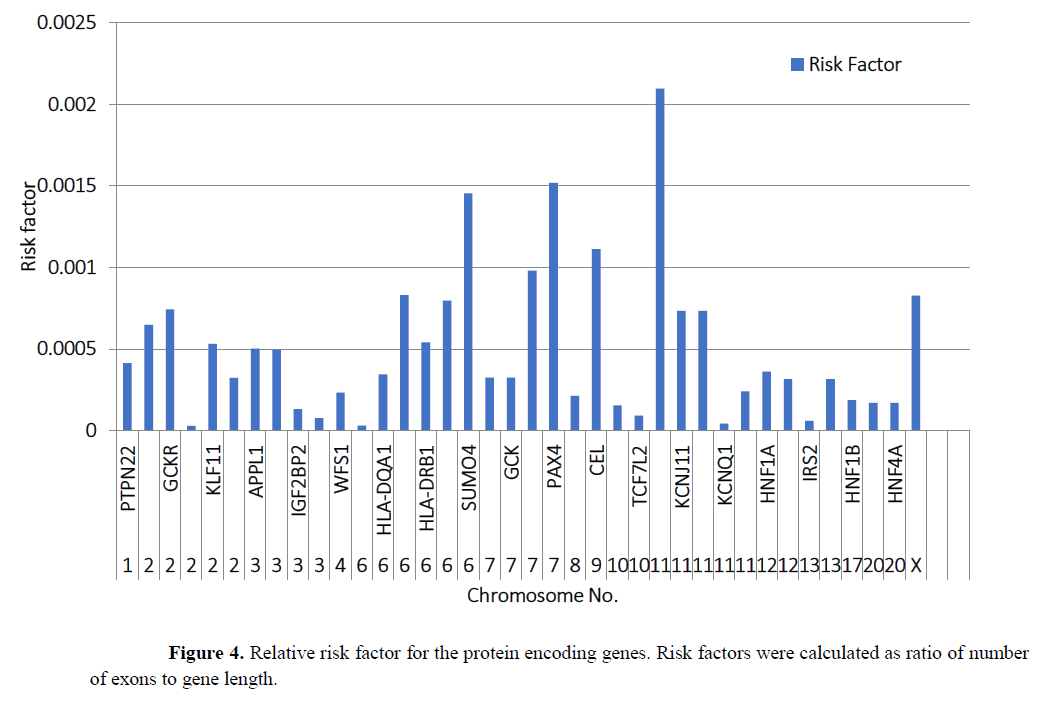
The risk of SNP is directly proportional to the number of exons as the risk of exposure for mutations is high. The risk of SNP is indirectly proportional to the length of the gene, since greater length with higher number of introns poses lesser risk of mutation in the protein encoded. Therefore, the risk for the protein encoding genes was derived as ratio of the number of exons to the length of the gene. It was observed that INS gene located on the chromosome 11 has greater susceptibility to mutation with the risk ratio value of 0.0020, followed by PAX4 gene located on chromosome 7 with a value of 0.0015 and SUMO4 gene located on chromosome 6 with a value of 0.0014. The relative risk for the rest of the genes was presented in Figure 4. It can be noted that INS gene has direct relevance to diabetes as the gene encodes the hormone insulin which facilitates the utilization of glucose by the peripheral tissues thus reducing the blood glucose level. Mutation in INS gene may lead to elevated blood glucose level.
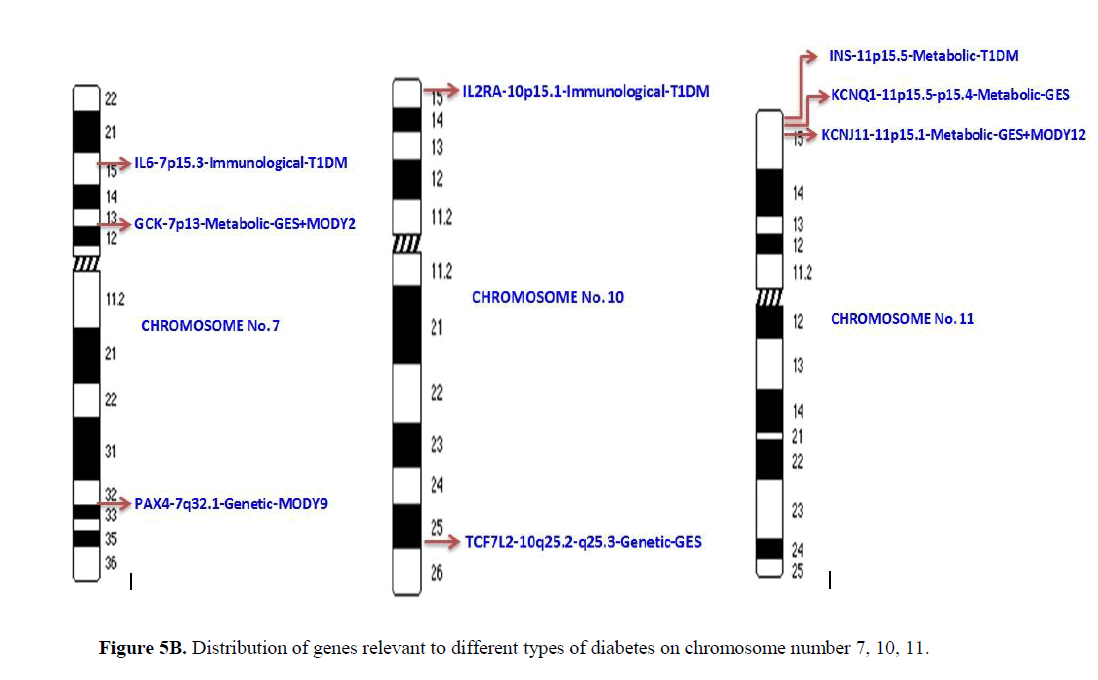
The distribution of the genes which are relevant to different types of diabetes on different chromosomes is presented in Figures 5A-5C. Only two or more genes present on single chromosome were represented. Chromosome number 1, 4, 8, 9, 17 and X harbored only single risk gene. Chromosome number 2 has 5 genes with metabolic, immunological and genetic effects. Chromosome number 3 has 4 genes with all the three categories of effects. Chromosome 6 has 6 genes with all three functional effects. Similarly, chromosome number 7 has 4 genes with all three functional effects. Chromosome number 10 has 3 genes with genetic and immunological effects. It is interesting to note that chromosome number 11 has 5 genes, all with metabolic relevance only. Chromosome 12, 13 and 20 have 2 genes each with common genetic effects. It is important to note that IRS1 gene located on chromosome number 2 is a risk factor for both T1DM and gestational diabetes, while HNF1A gene located on chromosome number 12 is a genetic risk factor for T1DM, T2DM and MODY3.
Epigenetic regulation of diabetic condition
Several dietary and environmental factors, both biotic and abiotic have been implicated in the epigenetic modifications. Viral infections and toxins also alter the routine metabolism that may predispose for epigenetic changes. These changes affect the genes by inducing either allele polymorphism mediated susceptibility or resistance to disease conditions (Desiderio et al., 2016; Zullo et al., 2017). Metabolic compounds such as S-adenosylmethionine, acetyl-CoA, nicotinamide adenine dinucleotide, α-ketoglutarate, and adenosine triphosphate function as cofactors for methyltransferases, deacetylases and kinases which modify chromatin. Dietary components such as free fatty acids, amino acids, glucose, glutamine, etc., modulate epigenetics (Etchegaray & Mostoslavsky, 2016). Aberrant toll-like receptor (TLR) signaling causes autoimmune response leading to development of T1DM. Epigenetic changes alter TLR signaling leading to T1DM (Xie et al., 2018). BCG vaccination was found to reduce long term hyperglycemia in T1DM patients. BCG vaccination was found to epigenetically reset T regulatory genes with genetic reprogramming of tolerance and induce aerobic glycolysis resulting in glucose homeostasis (Kuhtreiber et al., 2018). MicroRNAs are small non-coding RNA that regulates gene expression. During the onset of T1DM, overexpression of miR-487a-3p was noted that repress CTLA4 and FOXO3 by binding to their 3'UTRs and thus resulting in manifestation of T1DM (Zurawek et al., 2018).
DNA methylation has been found as a key mechanism for instigation of T2DM affecting both insulin secretion as well as insulin sensitivity. Based on a review by Zhou et al. have described 17 different genes that are involved in epigenetic manifestation of T2DM (Zhou, Sun, Li, & Zhu, 2018). Out of them two genes (insulin and metformin transporter genes) in islet and liver respectively were found susceptible to demethylation. Four out of the seventeen genes (LINE-1, MCP-1, TXNIP, NR4A1) in peripheral blood were susceptible to hypomethylation. PPARGC1A, KCNQ1, Insulin gene, PDX-1, GLP1R, MEG3-DLK1 microRNA, NDUFB6, COX7A1, IGFBP1 and IGFBP7, Alu repeats, TCF7L2 belonging to different cell types were susceptible to hypermethylation. Insulin gene, PPARGC1A, PDX-1, MEG3-DLK1 and microRNA were associated with insulin secretion. NDUFB6, COX7A1, IGFBP1, IGFBP7 and Alu repeats were associated with insulin sensitivity. Metformin transporter genes, KCNQ1, Insulin gene, GLP1R, TCF7L2, LINE-1, MCP-1, TXNIP and NR4A1 were associated with glycemic metabolism. Methylation of mitochondrial DNA has been regarded as the most prominent in microvascular complication of T2DM (Rorbach-Dolata, Kubis, & Piwowar, 2017). Acetylation of arginine and lysine among histone proteins was observed as an important epigenetic modification, key to the development of T2DM.
Genetic challenges, success and future directions in mitigating diabetic condition
Recently, GWAS studies revealed over 60 susceptible genes marked with SNPs that participate in gene regulatory network in manifestation of T1DM (Nyaga, Vickers, Jefferies, Perry, & O’Sullivan, 2018). About 80% of the genetic heritability of T1DM can be explained with GWAS data (Pociot et al., 2010). However, most of the associated variants are present in the non-coding region. Within immune system domain, about 56 genetic risk variants were found to be associated with T1DM out of which 11 were shared with endothelial functioning and 2 were shared with islet functioning. Endothelial and pancreatic islets have 2 exclusive genetic risk factors each with relevance to T1DM (Wallet, Santostefano, Terada, & Brusko, 2017). The risk associated with non-immunogenic genes is relatively less. However, a large number of variants with low frequency may influence the autoimmune functioning and thus the manifestation of T1DM. One such risk locus is UBASH3A gene located on chromosome 21q22.3. that inhibits NF-κB signaling (Ge & Concannon, 2018). In the MHC region, rs9260151 and rs3135002 have been linked with the risk of T1DM (Roshandel et al., 2018). In a different study, Erb-b2 receptor tyrosine kinase 3 (ERBB3) rs2292239 SNP was found to be associated with risk for T1DM in a white Brazilian population and has been identified as a susceptibility locus for T1DM (Lemos et al., 2018). Zhu et al. have emphasized that induction of immune tolerance together with genetic engineering could be beneficial in islet transplantation in pancreas as cellular replacement therapy for T1DM (Zhu et al., 2018).
It was found that monogenic forms of diabetes such as MODY can be treated with good precision. However, it was found that the prevalence of monogenic diabetes was found to be different across different geographical regions and demographics. Therefore, it is important to widely characterize the genetic basis of monogenic diabetes to accurately differentiate from other forms of adult onset diabetes. Intolerance towards β-cell antigen is the main reason for instigation of T1DM. For this, antigen specific immunotherapy confers immune tolerance either by supply of specific antigens that are tolerated or by promoting tolerance responses. Creusot et al. have identified that alteration in the functioning of the antigen presenting cells in T1DM as affected by genetic polymorphism is a challenge for antigen specific immunotherapy of T1DM (Creusot, Postigo-Fernandez, & Teteloshvili, 2018). The study emphasized on better characterization of such alterations for devising efficient methods for antigen tolerance.
Meta-analysis considering multiple factors is essential for understanding the associations of the genes by combining different studies. Case control ratio influences the power and reliability of the outcome. Yang et al. proposed a novel approach by incorporating adjustments for population stratification by correcting for known population structure through minor allele frequencies and emphasized its applicability for both single gene and gene level association studies (Yang, Chen, & Abecasis, 2018).
Conclusion
Epigenetic factors and non-immunogenic genetic factors seem to pose lower risk when compared to immunogenic genetic determinants in the manifestation of T1DM. It remains to be inferred, whether the genetic susceptibility or the risk factors associated with autoimmune mechanism is more prominent in the manifestation of T1DM than epigenetic causes. The probability of effective genetic therapy is high for single gene involvement when compared to polygenic involvement. Particularly, T1DM has involvement of multiple genes with immunological role. Therefore, identification of specific genetic targets in polygenic cases seems to be ineffective. However, T1DM is associated with insulin secretion which is a function of single type of cells. Therefore, the study of genes associated with islet functioning such as SLC30A8, DDX1, SRp55 would bring in certain degree of specificity to preserve and sustain β-cells and their insulin secretion potential. The factors that preserve the beta cell functioning, suppression of autoimmune response towards islets need to be identified and characterized for effective prevention of T1DM. As there are susceptible alleles, there are also protective variants of genes associated with T1DM such as A/G splice-site single nucleotide polymorphism (SNP; rs10774671) in OAS1 gene. Further exploration of beneficial polymorphisms associated with insulin secretion and insulin sensitivity need to be carried out. Epigenetic changes have role in instigation of diabetes; hence, epigenetic drugs may have substantial effect on the prevention and treatment of T1DM. Manifestation of T2DM is associated with multiple genes. Compared to other forms of diabetes, the genetic risk factors with genetic and metabolic functioning for the manifestation of T2DM are Epigenetic factors and non-immunogenic genetic factors seem to pose lower risk when compared to immunogenic genetic determinants in the manifestation of T1DM. It remains to be inferred, whether the genetic susceptibility or the risk factors associated with autoimmune mechanism is more prominent in the manifestation of T1DM than epigenetic causes. The probability of effective genetic therapy is high for single gene involvement when compared to polygenic involvement. Particularly, T1DM has involvement of multiple genes with immunological role. Therefore, identification of specific genetic targets in polygenic cases seems to be ineffective. However, T1DM is associated with insulin secretion which is a function of single type of cells. Therefore, the study of genes associated with islet functioning such as SLC30A8, DDX1, SRp55 would bring in certain degree of specificity to preserve and sustain β-cells and their insulin secretion potential. The factors that preserve the beta cell functioning, suppression of autoimmune response towards islets need to be identified and characterized for effective prevention of T1DM. As there are susceptible alleles, there are also protective variants of genes associated with T1DM such as A/G splice-site single nucleotide polymorphism (SNP; rs10774671) in OAS1 gene. Further exploration of beneficial polymorphisms associated with insulin secretion and insulin sensitivity need to be carried out. Epigenetic changes have role in instigation of diabetes; hence, epigenetic drugs may have substantial effect on the prevention and treatment of T1DM. Manifestation of T2DM is associated with multiple genes. Compared to other forms of diabetes, the genetic risk factors with genetic and metabolic functioning for the manifestation of T2DM are
About the Authors
Corresponding Author
Dhilleswara Rao Vana
Center for Excellence in Genomics, Madhurai Kamaraj University, Tamil Nadu, India
- Email:
- drvana.biotech@gmail.com
Dattatreya Adapa
Alumnus, Department of Microbiology, Food Science and Technology, GITAM University, Andhra Pradesh, India
- Email:
- dattu619@gmail.com
References
- Agostini M, Schoenmakers E, Beig J, Fairall L, et al. (2018) A pharmacogenetic approach to the treatment of patients with PPARG mutations. Diabetes 67: 1086-1092. https://doi.org/10.2337/db17-1236
- Aly TA, Ide A, Jahromi MM, Barker JM, et al. (2006) Extreme genetic risk for type 1A diabetes. Proceedings of the National Academy of Sciences 103: 14074-14079. https://doi.org/10.1073/pnas.0606349103
- American Diabetes Association (2017) Classification and diagnosis of diabetes. Diabetes Care 40: S11-S24. https://doi.org/10.2337/dc17-s005
- Arechiga AF, Habib T, He Y, Zhang X, et al. (2009) Cutting edge: the PTPN22 allelic variant associated with autoimmunity impairs B cell signaling. The Journal of Immunology 182: 3343-3347. https://doi.org/10.4049/jimmunol.0713370
- Arunagiri A, Haataja L, Cunningham CN, Shrestha N, et al. (2018) Misfolded proinsulin in the endoplasmic reticulum during development of beta cell failure in diabetes. Annals of the New York Academy of Sciences 1418: 5-19. https://doi.org/10.1111/nyas.13531
- Atkinson MA, Eisenbarth GS, Michels AW (2014) Type 1 diabetes. Lancet (London, England), 383: 69-82. https://doi.org/10.1016/s0140-6736 (13)60591-7
- Ay A, Alkanli N, Sipahi T, Gulyasar T, et al. (2018) Investigation of the relationship between MTHFR, IRS and CALCA gene polymorphisms and development of diabetic nephropathy in patients with type 2 diabetes mellitus. Biotechnology & Biotechnological Equipment 1-9. https://doi.org/10.1080/13102818.2018.1485514
- Bansal V, Boehm BO, Darvasi A (2018) Identification of a missense variant in the WFS1 gene that causes a mild form of Wolfram syndrome and is associated with risk for type 2 diabetes in Ashkenazi Jewish individuals. Diabetologia 61: 2180-2188. https://doi.org/10.1007/s00125-018-4690-3
- Barbetti F and D’Annunzio G (2018) Genetic causes and treatment of neonatal diabetes and early childhood diabetes. Best Practice & Research. Clinical Endocrinology & Metabolism 32: 575-591. https://doi.org/10.1016/j.beem.2018.06.008
- Barbetti F, Mammì C, Liu M, Grasso V, et al. (2017) Neonatal Diabetes: Permanent Neonatal Diabetes and Transient Neonatal Diabetes. In Diabetes Associated with Single Gene Defects and Chromosomal Abnormalities. Karger Publishers, 25: 1-25. https://doi.org/10.1159/000454748
- Barnett AH, Eff C, Leslie RDG, Pyke DA (1981) Diabetes in identical twins. Diabetologia 20: 87-93. https://doi.org/10.1007/bf00262007
- Barrett JC, Clayton DG, Concannon P, Akolkar B, et al. (2009) Genome-wide association study and meta-analysis find that over 40 loci affect risk of type 1 diabetes. Nature Genetics 41: 703-707. https://doi.org/10.1038/ng.381
- Baschal EE and Eisenbarth GS (2008) Extreme genetic risk for type 1A diabetes in the post-genome era. Journal of Autoimmunity 31: 1-6. https://doi.org/10.1016/j.jaut.2008.03.003
- Ben Khelifa S, Martinez R, Dandana A, Khochtali I, et al. (2018) Maturity Onset Diabetes of the Young (MODY) in Tunisia: Low frequencies of GCK and HNF1A mutations. Gene 651: 44-48. https://doi.org/10.1016/j.gene.2018.01.081
- Berndt SI, Gustafsson S, Magi R, Ganna A, et al. (2013) Genome-wide meta-analysis identifies 11 new loci for anthropometric traits and provides insights into genetic architecture. Nature Genetics 45: 501-512. https://doi.org/10.1038/ng.2606
- Biesecker LG (2018) Genomic screening for monogenic forms of diabetes. BMC Medicine England 16. https://doi.org/10.1186/s12916-018-1012-z
- Bonnefond A, Philippe J, Durand E, Muller J, et al. (2014) Highly sensitive diagnosis of 43 monogenic forms of diabetes or obesity through one-step PCR-based enrichment in combination with next-generation sequencing. Diabetes Care 37: 460-467. https://doi.org/10.2337/dc13-0698
- Brownlie RJ, Zamoyska R, Salmond RJ (2018) Regulation of autoimmune and anti-tumour T-cell responses by PTPN22. Immunology 154: 377-382. https://doi.org/10.1111/imm.12919
- Caillat-Zucman S (2017) New insights into the understanding of MHC associations with immune-mediated disorders. HLA 89: 3-13. https://doi.org/10.1111/tan.12947
- Catalano PM (2014) Trying to understand gestational diabetes. Diabetic Medicine : A Journal of the British Diabetic Association 31: 273-281. https://doi.org/10.1111/dme.12381
- Chen YL, Qiao YC, Pan YH, Xu Y, et al. (2017) Correlation between serum interleukin-6 level and type 1 diabetes mellitus: A systematic review and meta-analysis. Cytokine 94: 14-20. https://doi.org/10.1016/j.cyto.2017.01.002
- Claussnitzer M, Dankel SN, Klocke B, Grallert H, et al. (2014) Leveraging cross-species transcription factor binding site patterns: from diabetes risk loci to disease mechanisms. Cell 156: 343-358.
- Cooper DN, Krawczak M, Polychronakos C, Tyler-Smith C, et al. (2013) Where genotype is not predictive of phenotype: towards an understanding of the molecular basis of reduced penetrance in human inherited disease. Human Genetics 132: 1077-1130. https://doi.org/10.1007/s00439-013-1331-2
- Copps KD and White MF (2012) Regulation of insulin sensitivity by serine/threonine phosphorylation of insulin receptor substrate proteins IRS1 and IRS2. Diabetologia 55: 2565-2582. https://doi.org/10.1007/s00125-012-2644-8
- Costa CD, Teleginski A, Al-Lahham Y, Souza EM, et al. (2018) Association of Polymorphism rs17576 of the Metalloproteinase 9 Gene with Gestational Diabetes in Euro-Brazilian Pregnant Women. Clinical Laboratory 64: 645-649. https://doi.org/10.7754/clin.lab.2017.171125
- Creusot RJ, Postigo-Fernandez J, Teteloshvili N (2018) Altered Function of Antigen-Presenting Cells in Type 1 Diabetes: A Challenge for Antigen-Specific Immunotherapy? Diabetes 67: 1481-1494. https://doi.org/10.2337/db17-1564
- Damm P (1998) Gestational diabetes mellitus and subsequent development of overt diabetes mellitus. Danish Medical Bulletin 45: 495-509.
- De Rosa S, Arcidiacono B, Chiefari E, Brunetti A, et al. (2018) Type 2 Diabetes Mellitus and Cardiovascular Disease: Genetic and Epigenetic Links . Frontiers in Endocrinology 9. https://doi.org/10.3389/fendo.2018.00002
- Desiderio A, Spinelli R, Ciccarelli M, Nigro C, et al. (2016) Epigenetics: spotlight on type 2 diabetes and obesity. Journal of Endocrinological Investigation 39: 1095-1103. https://doi.org/10.1007/s40618-016-0473-1
- Dupuis J, Langenberg C, Prokopenko I, Saxena R, et al. (2010) New genetic loci implicated in fasting glucose homeostasis and their impact on type 2 diabetes risk. Nature Genetics 42:105-116. https://doi.org/10.1038/ng.520
- Ellard S, Bellanne-Chantelot C, Hattersley AT (2008) Best practice guidelines for the molecular genetic diagnosis of maturity-onset diabetes of the young. Diabetologia 51: 546–553. https://doi.org/10.1007/s00125-008-0942-y
- Ellard S, Lango Allen H, De Franco E, Flanagan SE, et al. (2013) Improved genetic testing for monogenic diabetes using targeted next-generation sequencing. Diabetologia 56: 1958-1963. https://doi.org/10.1007/s00125-013-2962-5
- Estrada K, Aukrust I, Bjørkhaug L, Burtt NP, et al. (2014) Association of a Low-Frequency Variant in HNF1A With Type 2 Diabetes in a Latino Population. Jama 311: 2305. https://doi.org/10.1001/jama.2014.6511
- Etchegaray JP and Mostoslavsky R (2016) Interplay between Metabolism and Epigenetics: A Nuclear Adaptation to Environmental Changes. Molecular Cell 62: 695-711. https://doi.org/10.1016/j.molcel.2016.05.029
- Field LL, Bonnevie-Nielsen V, Pociot F, Lu S (2005) OAS1 splice site polymorphism controlling antiviral enzyme activity influences susceptibility to type 1 diabetes. Diabetes 54: 1588-1591. https://doi.org/10.2337/diabetes.54.5.1588
- Flannick J, Beer NL, Bick AG, Agarwala V, et al. (2013) Assessing the phenotypic effects in the general population of rare variants in genes for a dominant Mendelian form of diabetes. Nature Genetics 45: 1380-1385. https://doi.org/10.1038/ng.2794
- Flannick J, Thorleifsson G, Beer NL, Jacobs SBR, et al. (2014) Loss-of-function mutations in SLC30A8 protect against type 2 diabetes. Nature Genetics 46: 357–363. https://doi.org/10.1038/ng.2915
- Florez JC (2008) Newly identified loci highlight beta cell dysfunction as a key cause of type 2 diabetes: Where are the insulin resistance genes? Diabetologia 51: 1100-1110. https://doi.org/10.1007/s00125-008-1025-9
- Florez JC, Jablonski KA, McAteer JB, Franks PW, et al. (2012) Effects of genetic variants previously associated with fasting glucose and insulin in the Diabetes Prevention Program. PloS One 7: e44424. https://doi.org/10.1371/journal.pone.0044424
- Fogarty MP, Cannon ME, Vadlamudi S, Gaulton KJ (2014) Identification of a regulatory variant that binds FOXA1 and FOXA2 at the CDC123/CAMK1D type 2 diabetes GWAS locus. PLoS Genetics 10: e1004633. https://doi.org/10.1371/journal.pgen.1004633
- Fogarty MP, Panhuis TM, Vadlamudi S, Buchkovich ML, et al. (2013) Allele-specific transcriptional activity at type 2 diabetes-associated single nucleotide polymorphisms in regions of pancreatic islet open chromatin at the JAZF1 locus Diabetes 62: 1756-1762. https://doi.org/10.2337/db12-0972
- Forbes JM, and Cooper ME (2013) Mechanisms of diabetic complications. Physiological Reviews 93: 137-188. https://doi.org/10.1152/physrev.00045.2011
- Franks PW, Rolandsson O, Debenham SL, Fawcett KA, et al. (2008) Replication of the association between variants in WFS1 and risk of type 2 diabetes in European populations. Diabetologia 51: 458-463. https://doi.org/10.1007/s00125-007-0887-6
- Franzago M, Fraticelli F, Marchetti D, Celentano C, et al. (2018) Nutrigenetic variants and cardio-metabolic risk in women with or without gestational diabetes. Diabetes Research and Clinical Practice 137: 64-71. https://doi.org/10.1016/j.diabres.2018.01.001
- Froguel P (2015) Obesity genes and their effect on the energy balance. Bulletin de l’Academie nationale de medecine 199: 1269-1279.
- García-Chapa EG, Leal-Ugarte E, Peralta-Leal V, Durán-González J, et al. (2017) Genetic Epidemiology of Type 2 Diabetes in Mexican Mestizos. BioMed Research International 2017: 1-10. https://doi.org/10.1155/2017/3937893
- Gardner DS and Tai ES (2012) Clinical features and treatment of maturity onset diabetes of the young (MODY). Diabetes, Metabolic Syndrome and Obesity : Targets and Therapy 5: 101-108. https://doi.org/10.2147/dmso.s23353
- Ge Y and Concannon P (2018) Molecular-genetic characterization of common, noncoding UBASH3A variants associated with type 1 diabetes. European Journal of Human Genetics 26: 1060-1064. https://doi.org/10.1038/s41431-018-0123-5
- Grubic Z, Putarek NR, Maskalan M, Zunec R, et al. (2018) Human Leukocyte Antigen class II polymorphisms among Croatian patients with type 1 diabetes and autoimmune polyglandular syndrome type 3 variant. Gene. 674: 93-97. https://doi.org/10.1016/j.gene.2018.06.083
- Hansen L and Pedersen O (2004) Genetics of type 2 diabetes mellitus: status and perspectives. Diabetes, Obesity and Metabolism 7: 122-135. https://doi.org/10.1111/j.1463-1326.2004.00396.x
- Hasanvand Z, Sadeghi A, Rezvanfar MR, Goodarzi MT, et al. (2018) Association between chemerin rs17173608 and rs4721 gene polymorphisms and gestational diabetes mellitus in Iranian pregnant women. Gene 649: 87-92. https://doi.org/10.1016/j.gene.2018.01.061
- Hashemzadeh AA, Nasoohi N, Raygan F, Aghadavod E, et al. (2017) Flaxseed Oil Supplementation Improve Gene Expression Levels of PPAR-gamma, LP (a), IL-1 and TNF-alpha in Type 2 Diabetic Patients with Coronary Heart Disease. Lipids 52: 907-915. https://doi.org/10.1007/s11745-017-4295-5
- Hofmann S, Philbrook C, Gerbitz KD, Bauer MF (2003) Wolfram syndrome: structural and functional analyses of mutant and wild-type wolframin, the WFS1 gene product . Human Molecular Genetics 12: 2003-2012. https://doi.org/10.1093/hmg/ddg214
- Holmkvist J, Cervin C, Lyssenko V, Winckler W, et al. (2006) Common variants in HNF-1 alpha and risk of type 2 diabetes. Diabetologia 49: 2882-2891. https://doi.org/10.1007/s00125-006-0450-x
- Huerta-Saenz L, Saunders C, Yan Y (2018) Challenging diagnosis of congenital hyperinsulinism in two infants of diabetic mothers with rare pathogenic KCNJ11 and HNF4A gene variants. International Journal of Pediatric Endocrinology 2018: 5. https://doi.org/10.1186/s13633-018-0060-7
- Hwang JL, Park S, Ye H, Sanyoura M, et al. (2018) FOXP3 mutations causing early‐onset insulin‐requiring diabetes but without other features of immune dysregulation, polyendocrinopathy, enteropathy, X‐linked syndrome. Pediatric Diabetes 19: 388-392. https://doi.org/10.1111/pedi.12612
- Ismail-Beigi F, Craven T, Banerji MA, Basile J, et al. (2010) Effect of intensive treatment of hyperglycaemia on microvascular outcomes in type 2 diabetes: an analysis of the ACCORD randomised trial. Lancet 376: 419-430. https://doi.org/10.1016/s0140-6736 (10)60576-4
- Jamalpour S, Zain SM, Mosavat M, Mohamed Z, et al. (2018) A case-control study and meta-analysis confirm glucokinase regulatory gene rs780094 is a risk factor for gestational diabetes mellitus. Gene 650: 34-40. https://doi.org/10.1016/j.gene.2018.01.091
- Jerram ST and Leslie RD (2017) The genetic architecture of type 1 diabetes. Genes 8: 209. https://doi.org/10.3390/genes8080209
- Johnson MB, Patel KA, De Franco E, Houghton JAL, et al. (2018) A type 1 diabetes genetic risk score can discriminate monogenic autoimmunity with diabetes from early-onset clustering of polygenic autoimmunity with diabetes Diabetologia 61: 862-869. https://doi.org/10.1007/s00125-018-4551-0
- Juan-Mateu J, Alvelos MI, Turatsinze JV, Villate O, et al. (2018) SRp55 Regulates a Splicing Network That Controls Human Pancreatic beta-Cell Function and Survival. Diabetes 67: 423-436. https://doi.org/10.2337/db17-0736
- Kang J, Lee CN, Li HY, Hsu KH, et al. (2017) Genome-wide DNA methylation variation in maternal and cord blood of gestational diabetes population. Diabetes Research and Clinical Practice 132: 127-136. https://doi.org/10.1016/j.diabres.2017.07.034
- Kawai VK, Levinson RT, Adefurin A, Kurnik D, et al. (2017) Variation in the alpha2A-adrenergic receptor gene and risk of gestational diabetes. Pharmacogenomics 18: 1381-1386. https://doi.org/10.2217/pgs-2017-0079
- Kerru N, Singh-Pillay A, Awolade P, Singh P (2018) Current anti-diabetic agents and their molecular targets: A review. European Journal of Medicinal Chemistry 152: 436-488. https://doi.org/10.1016/j.ejmech.2018.04.061
- Kleiner S, Gomez D, Megra B, Na E, et al. (2018) Mice harboring the human SLC30A8 R138X loss-of-function mutation have increased insulin secretory capacity. Proceedings of the National Academy of Sciences of the United States of America 115: E7642–E7649. https://doi.org/10.1073/pnas.1721418115
- Kooner JS, Saleheen D, Sim X, Sehmi J, et al. (2011) Genome-wide association study in individuals of South Asian ancestry identifies six new type 2 diabetes susceptibility loci. Nature Genetics 43: 984–989 https://doi.org/10.1038/ng.921
- Kota SK, Meher LK, Jammula S, Kota SK, et al. (2012) Genetics of type 2 diabetes mellitus and other specific types of diabetes; its role in treatment modalities. Diabetes & Metabolic Syndrome 6: 54-58. https://doi.org/10.1016/j.dsx.2012.05.014
- Kuhtreiber WM, Tran L, Kim T, Dybala M, et al. (2018) Long-term reduction in hyperglycemia in advanced type 1 diabetes: the id of induced aerobic glycolysis with BCG vaccinations. NPJ Vaccines 3: 23. https://doi.org/10.1038/s41541-018-0062-8
- Lecarpentier Y, Claes V, Vallée A, Hébert JL (2017) Interactions between PPAR Gamma and the Canonical Wnt/Beta-Catenin Pathway in Type 2 Diabetes and Colon Cancer. PPAR Research 2017: 1-9. https://doi.org/10.1155/2017/5879090
- Lemos NE, Dieter C, Dorfman LE, Assmann TS, et al. (2018) The rs2292239 polymorphism in ERBB3 gene is associated with risk for type 1 diabetes mellitus in a Brazilian population. Gene 644: 122-128. https://doi.org/10.1016/j.gene.2017.11.009
- Li M, Liu J, Yi H, Xu L, et al. (2018) A novel mutation of WFS1 gene in a Chinese patient with Wolfram syndrome: a case report. BMC Pediatrics 18: 116. https://doi.org/10.1186/s12887-018-1091-1
- Liu Y, Tan Q, Liu F (2017) Differentially methylated circulating DNA: A novel biomarker to monitor beta cell death. Journal of Diabetes and Its Complications 32: 349–353. https://doi.org/10.1016/j.jdiacomp.2017.08.012
- Locke AE, Kahali B, Berndt SI, Justice AE, et al. (2015) Genetic studies of body mass index yield new insights for obesity biology. Nature 518: 197-206. https://doi.org/10.1038/nature14177
- López Rodríguez M, Kaminska D, Lappalainen K, Pihlajamäki J, et al. (2017) Identification and characterization of a FOXA2-regulated transcriptional enhancer at a type 2 diabetes intronic locus that controls GCKR expression in liver cells. Genome Medicine 9: 63. https://doi.org/10.1186/s13073-017-0453-x
- Majithia AR, Flannick J, Shahinian P, Guo M, et al. (2014) Rare variants in PPARG with decreased activity in adipocyte differentiation are associated with increased risk of type 2 diabetes. Proceedings of the National Academy of Sciences of the United States of America 111: 13127-13132. https://doi.org/10.1073/pnas.1419356111
- Manning AK, Hivert MF, Scott RA, Grimsby JL, et al. (2012) A genome-wide approach accounting for body mass index identifies genetic variants influencing fasting glycemic traits and insulin resistance. Nature Genetics 44: 659-669. https://doi.org/10.1038/ng.2274
- Manu MS, Rachana KS, Advirao GM (2017) Altered expression of IRS2 and GRB2 in demyelination of peripheral neurons: Implications in diabetic neuropathy. Neuropeptides 62: 71-79. https://doi.org/10.1016/j.npep.2016.12.004
- Mathers CD and Loncar D (2006) Projections of global mortality and burden of disease from 2002 to 2030. PLoS Medicine 3: e442. https://doi.org/10.1371/journal.pmed.0030442
- McCarthy MI (2004) Progress in defining the molecular basis of type 2 diabetes mellitus through susceptibility-gene identification. Human Molecular Genetics 13: 33R-41. https://doi.org/10.1093/hmg/ddh057
- McCarthy MI (2010) Genomics, type 2 diabetes, and obesity. New England Journal of Medicine 363: 2339-2350. https://doi.org/10.1056/nejmra0906948
- Minton JAL, Hattersley AT, Owen K, McCarthy MI, et al. (2002) Association studies of genetic variation in the WFS1 gene and type 2 diabetes in UK populations. Diabetes 51: 1287-1290. https://doi.org/10.2337/diabetes.51.4.1287
- Molven A and Njolstad PR (2011) Role of molecular genetics in transforming diagnosis of diabetes mellitus. Expert Review of Molecular Diagnostics 11: 313-320. https://doi.org/10.1586/erm.10.123
- Morris AP, Voight BF, Teslovich TM, Ferreira T, et al. (2012) Large-scale association analysis provides insights into the genetic architecture and pathophysiology of type 2 diabetes. Nature Genetics 44: 981-990. https://doi.org/10.1038/ng.2383
- NCD-RisC NRFC (2016) Worldwide trends in diabetes since 1980: a pooled analysis of 751 population-based studies with 4.4 million participants. Lancet 387: 1513-1530. https://doi.org/10.1016/s0140-6736 (16)00618-8
- Noble JA and Valdes AM (2011) Genetics of the HLA region in the prediction of type 1 diabetes. Current Diabetes Reports 11: 533-542. https://doi.org/10.1007/s11892-011-0223-x
- Nuli R, Azati J, Shawuti Z, Shan J, et al. (2018) The relationship between HNF1A variations and type 2 diabetes mellitus risk factors in the Uyghur population in Urumqi. International Journal of Clinical And Experimental Medicine 11: 957-965.
- Nyaga DM, Vickers MH, Jefferies C, Perry JK, et al. (2018) The genetic architecture of type 1 diabetes mellitus. Molecular and Cellular Endocrinology. 477: 70-80. https://doi.org/10.1016/j.mce.2018.06.002
- Olefsky, J. M. (2000). Treatment of insulin resistance with peroxisome proliferator–activated receptor γ agonists. The Journal of Clinical Investigation 106: 467-472. https://doi.org/10.1172/jci10843
- Ono K, Igata M, Kondo T, Kitano S, et al. (2018) Identification of microRNA that represses IRS-1 expression in liver. PLOS ONE 13: e0191553. https://doi.org/10.1371/journal.pone.0191553
- Pace NP, Craus J, Felice A, Vassallo J (2018) Case Report: Identification of an HNF1B p.Arg527Gln mutation in a Maltese patient with atypical early onset diabetes and diabetic nephropathy. BMC Endocrine Disorders 18: 28. https://doi.org/10.1186/s12902-018-0257-z
- Pavic T, Juszczak A, Pape Medvidovic E, Burrows C, et al. (2018) Maturity onset diabetes of the young due to HNF1A variants in Croatia. Biochemia Medica 28: 20703. https://doi.org/10.11613/bm.2018.020703
- Pierce M, Keen H, Bradley C (1995) Risk of Diabetes in Offspring of Parents with Non‐insulin‐dependent Diabetes. Diabetic Medicine 12: 6-13. https://doi.org/10.1111/j.1464-5491.1995.tb02054.x
- Pociot F, Akolkar B, Concannon P, Erlich HA, et al. (2010) Genetics of type 1 diabetes: What’s next? Diabetes. 59 issue 7: 1561-1571.https://doi.org/10.2337/db10-0076
- Pollak CF, Lagos LM, Santos MJL, Poggi H, et al. (2017) Diabetes mellitus caused by a mutation of glucokinase gene. Revista medica de Chile 145: 1203-1207. https://doi.org/10.4067/s0034-98872017000901203
- Rama Chandran S, Bhalshankar J, Farhad Vasanwala R, Zhao Y, et al. (2018) Traditional clinical criteria outperform high-sensitivity C-reactive protein for the screening of hepatic nuclear factor 1 alpha maturity-onset diabetes of the young among young Asians with diabetes. Therapeutic Advances in Endocrinology and Metabolism 9: 271-282. https://doi.org/10.1177/2042018818776167
- Randall JC, Winkler TW, Kutalik Z, Berndt SI, et al. (2013) Sex-stratified genome-wide association studies including 270,000 individuals show sexual dimorphism in genetic loci for anthropometric traits. PLoS Genetics, 9: e1003500. https://doi.org/10.1371/journal.pgen.1003500
- Redondo MJ, Steck AK, Pugliese A (2018) Genetics of type 1 diabetes. Pediatric Diabetes 19: 346-353. https://doi.org/10.1111/pedi.12597
- Rehman K, Akash MSH, Liaqat A, Kamal S, et al. (2017) Role of interleukin-6 in development of insulin resistance and type 2 diabetes mellitus. Critical ReviewsTM in Eukaryotic Gene Expression 27: 229-236. https://doi.org/10.1615/critreveukaryotgeneexpr.2017019712
- Rich SS (2006) Genetics of Diabetes and Its Complications. Journal of the American Society of Nephrology 17: 353 LP-360. https://doi.org/10.1681/asn.2005070770
- Rigoli L, Bramanti P, Di Bella C, De Luca F (2018) Genetic and clinical aspects of Wolfram syndrome 1, a severe neurodegenerative disease. Pediatric Research 83: 921. https://doi.org/10.1038/pr.2018.17
- Rorbach-Dolata A, Kubis A, Piwowar A (2017) Epigenetic modifications: An important mechanism in diabetic disturbances. Postepy Higieny i Medycyny Doswiadczalnej 71: 960-974. https://doi.org/10.5604/01.3001.0010.6156
- Rorsman P and Ashcroft FM (2018) Pancreatic β-Cell Electrical Activity and Insulin Secretion: Of Mice and Men. Physiological Reviews 98: 117-214. https://doi.org/10.1152/physrev.00008.2017
- Roshandel D, Gubitosi-Klug R, Bull SB, Canty AJ, et al. (2018) Meta-genome-wide association studies identify a locus on chromosome 1 and multiple variants in the MHC region for serum C-peptide in type 1 diabetes. Diabetologia 61: 1098-1111. https://doi.org/10.1007/s00125-018-4555-9
- Rubio-Cabezas O, Hattersley AT, Njolstad PR, Mlynarski W, et al. (2014) ISPAD Clinical Practice Consensus Guidelines 2014. The diagnosis and management of monogenic diabetes in children and adolescents. Pediatric Diabetes 15: 47-64. https://doi.org/10.1111/pedi.12772
- Sanyoura M, Philipson LH, Naylor R (2018) Monogenic Diabetes in Children and Adolescents : Recognition and Treatment Options. Current Diabetes Reports 18: 58. https://doi.org/10.1007/s11892-018-1024-2
- Saxena R, Hivert MF, Langenberg C, Tanaka T, et al. (2010) Genetic variation in GIPR influences the glucose and insulin responses to an oral glucose challenge. Nature Genetics 42: 142. https://doi.org/10.1038/ng.521
- Schallenberg S and Kretschmer K (2018) New insight into type 1 diabetes development: resolving early diabetogenic CD4+ T cell responses that precede seroconversion. Annals of Translational Medicine 6: 58-58. https://doi.org/10.21037/atm.2017.12.14
- Schwitzgebel VM (2014) Many faces of monogenic diabetes. Journal of Diabetes Investigation 5: 121-133. https://doi.org/10.1111/jdi.12197
- Scott RA, Lagou V, Welch RP, Wheeler E, et al. (2012) Large-scale association analyses identify new loci influencing glycemic traits and provide insight into the underlying biological pathways. Nature Genetics 44: 991. https://doi.org/10.1038/ng.2385
- Shahi A, Prasad V, Imam SS, Abdul MM, et al. (2018) Pathophysiological Ramifications of Diabetic Condition: A Review. Asian Journal of Biomedical and Pharmaceutical Sciences 8: 28-38. https://doi.org/10.4066/2249-622x.65.18-845
- Shungin D, Winkler TW, Croteau-Chonka DC, Ferreira T, et al. (2015) New genetic loci link adipose and insulin biology to body fat distribution. Nature 518: 187-196.
- Sladek R, Rocheleau G, Rung J, Dina C, et al. (2007). A genome-wide association study identifies novel risk loci for type 2 diabetes. Nature 445: 881- 885. https://doi.org/10.1038/nature05616
- Smyth DJ, Cooper JD, Lowe CE, Nutland S, et al. (2006) No evidence for association of OAS1 with type 1 diabetes in unaffected siblings or type 1 diabetic cases. Diabetes 55: 1525-1528. https://doi.org/10.2337/db05-1452
- Steinthorsdottir V, Thorleifsson G, Sulem P, Helgason H, et al. (2014, March). Identification of low-frequency and rare sequence variants associated with elevated or reduced risk of type 2 diabetes. Nature Genetics. 46: 294-298. https://doi.org/10.1038/ng.2882
- Stumvoll M, Goldstein BJ, van Haeften TW (2005) Type 2 diabetes: principles of pathogenesis and therapy. The Lancet 365: 1333-1346. https://doi.org/10.1016/s0140-6736 (05)61032-x
- Tagoma A, Alnek K, Kirss A, Uibo R, et al. (2018) MicroRNA profiling of second trimester maternal plasma shows upregulation of miR-195-5p in patients with gestational diabetes. Gene 672: 137-142. https://doi.org/10.1016/j.gene.2018.06.004
- Tallapragada DSP, Bhaskar S, Chandak GR (2015) New insights from monogenic diabetes for “common” type 2 diabetes. Frontiers in Genetics 6. https://doi.org/10.3389/fgene.2015.00251
- Tarnowski M, Wieczorek A, Dziedziejko V, Safranow K, et al. (2017) IL16 and IL18 gene polymorphisms in women with gestational diabetes. Ginekologia Polska 88: 249-254. https://doi.org/10.5603/gp.a2017.0047
- Tattersal RB, Fajans SS (1975) Prevalence of diabetes and glucose intolerance in 199 offspring of thirty-seven conjugal diabetic parents. Diabetes 24: 452-462. https://doi.org/10.2337/diabetes.24.5.452
- Tessier MC, Qu HQ, Fréchette R, Bacot F, et al. (2006) Type 1 diabetes and the OAS gene cluster: association with splicing polymorphism or haplotype? Journal of Medical Genetics 43: 129-132. https://doi.org/10.1136/jmg.2005.035212
- Thomas CC, Philipson LH (2015) Update on diabetes classification. The Medical Clinics of North America 99: 1-16. https://doi.org/10.1016/j.mcna.2014.08.015
- Udler MS, Kim J, von Grotthuss M, Bonas-Guarch S, et al. (2018) Clustering of Type 2 Diabetes Genetic Loci by Multi-Trait Associations Identifies Disease Mechanisms and Subtypes. BioRxiv. https://doi.org/10.1101/319509
- Voight BF, Scott LJ, Steinthorsdottir V, Morris AP, et al. (2010) Twelve type 2 diabetes susceptibility loci identified through large-scale association analysis. Nature Genetics 42: 579. https://doi.org/10.1038/ng.609
- Wallet MA, Santostefano KE, Terada N, Brusko TM (2017) Isogenic cellular systems model the impact of genetic risk variants in the pathogenesis of type 1 diabetes. Frontiers in Endocrinology 8: 276. https://doi.org/10.3389/fendo.2017.00276
- Wang Z, Long H, Chang C, Zhao M, et al. (2018) Crosstalk between metabolism and epigenetic modifications in autoimmune diseases: a comprehensive overview. Cellular and Molecular Life Sciences 75: 3353-3369 https://doi.org/10.1007/s00018-018-2864-2
- Wild S, Roglic G, Green A, Sicree R, et al. (2004) Global Prevalence of Diabetes: Estimates for the year 2000 and projections for 2030. Diabetes Care 27: 1047-1053https://doi.org/10.2337/diacare.27.5.1047
- Wiley F (2016) Monogenic Diabetes: Not Your “Typical” Diabetes. Diabetes Self-Management 33: 36-37.
- Xie Z, Huang G, Wang Z, Luo S, et al. (2018) Epigenetic regulation of Toll-like receptors and its roles in type 1 diabetes. Journal of Molecular Medicine 96 : 741-751. https://doi.org/10.1007/s00109-018-1660-7
- Xu T, Shi Y, Liu J, Liu Y, et al. (2018) The rs10229583 polymorphism near paired box gene 4 is associated with gestational diabetes mellitus in Chinese women. The Journal of International Medical Research 46: 115-121. https://doi.org/10.1177/0300060517714934
- Yan L, Feng J, Cheng F, Cui X, et al. (2018) Circular RNA expression profiles in placental villi from women with gestational diabetes mellitus. Biochemical and Biophysical Research Communications 498: 743-750. https://doi.org/10.1016/j.bbrc.2018.03.051
- Yang J, Chen S, Abecasis G (2018) Improved score statistics for meta-analysis in single-variant and gene-level association studies. Genetic Epidemiology 42: 333-343.
- Zhang J and Sun TQ (2009) IRS-2 and its involvement in diabetes and aging. Vitamins and Hormones 80: 389-407. https://doi.org/10.1016/s0083-6729 (08)00614-6
- Zheng L, Li C, Qi W H, Qiao BL, et al. (2017) mExpression of macrophage migration inhibitory factor gene in placenta tissue and its correlation with gestational diabetes mellitus. Zhonghua yi xue za zhi 97: 3388-3391.
- Zhong W, Li Z, Zhou M, Xu T, et al. (2018) DDX1 regulates alternative splicing and insulin secretion in pancreatic beta cells. Biochemical and Biophysical Research Communications 500: 751-757. https://doi.org/10.1016/j.bbrc.2018.04.147
- Zhou Z, Sun B, Li X, Zhu C (2018) DNA methylation landscapes in the pathogenesis of type 2 diabetes mellitus. Nutrition & Metabolism 15: 47. https://doi.org/10.1186/s12986-018-0283-x
- Zhu H, Zhang X, He Y, Yu L, et al. (2018) Research progress on the donor cell sources of pancreatic islet transplantation for treatment of diabetes mellitus. Chinese Journal of Reparative and Reconstructive Surgery 32: 104-111.
- Zoungas S, De Galan BE, Ninomiya T, Grobbee D, et al. (2009) Combined effects of routine blood pressure lowering and intensive glucose control on macrovascular and microvascular outcomes in patients with type 2 diabetes: New results from the ADVANCE trial. Diabetes Care 32: 2068-2074. https://doi.org/10.2337/dc09-0959
- Zullo A, Sommese L, Nicoletti G, Donatelli F, et al. (2017) Epigenetics and type 1 diabetes: mechanisms and translational applications. Translational Research : The Journal of Laboratory and Clinical Medicine 185: 85-93. https://doi.org/10.1016/j.trsl.2017.05.002
- Zurawek M, Dzikiewicz-Krawczyk A, Izykowska K, Ziolkowska-Suchanek I, et al. (2018) miR-487a-3p upregulated in type 1 diabetes targets CTLA4 and FOXO3. Diabetes Research and Clinical Practice 142: 146-153. https://doi.org/10.1016/j.diabres.2018.05.044
Download:
Full PDF- Share This