Glioblastoma in a 20-year-old female patient after Chikungunya infection
Received: October 29, 2018
Accepted: November 15, 2018
Published: January 05, 2019
Genet.Mol.Res. 18(1):
Introduction
Gliomas are tumors of the central nervous system, derived from progenitor glial cells, causing diffuse and infiltrative lesions. Among the diffuse gliomas, glioblastoma (GBM) represents the most malignant type. Recently the molecular information was introduced into the classification of the diffuse gliomas (Louis et al., 2016).It was estimated that 11,320 new cases of central nervous system tumors will occur in Brazil alone in 2018-2019(Instituto Nacional de Câncer, 2018). The discovery of viruses and genetic alterations has brought great contributions to the understanding of the pathways involved in glial oncogenesis.
CASE REPORT
A 20-year-old female student of African descent, without a family history of cancer and without exposure to toxic agents, born in the city of Rio de Janeiro where she resided, presented in April 2016 with an arboviral disease (cutaneous rash, artrlagia, and fever). A blood test was performed at the Pedro Ernesto Hospital (HUPE) of the State University of Rio de Janeiro (UERJ) and IgM+ was detected for the Chikungunya virus (CHIKV)
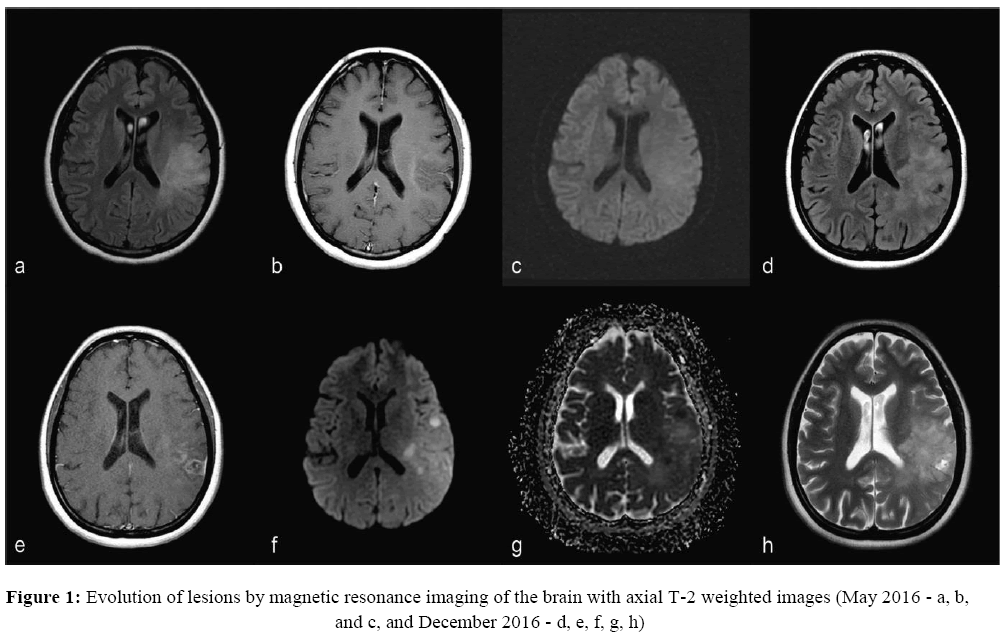
One month later, the patient presented with three seizures (two focal motor in the right hemiface and one generalized tonic-clonic). Magnetic resonance imaging (MRI), including an angiogram (MRA) excluded any lesions. In the following month cranial magnetic resonance imaging was performed, which detected a non-specific lesion, hyperintense in FLAIR/T2 in the front parietal region of the left hemisphere, without restricted diffusion (DWI) and without contrast enhancement (Figure 1). By this time she was taking Phenytoin 300 mg per day, Phenobarbital 150 mg per day, Oxcarbazepine 900 mg per day, and Clonazepam 0.5 mg 8/8 hours.
Notes:
1a - FLAIR shows expansive poorly defined lesion at left cortical -subcortical fronto-parietal region, extending to deep white matter and associated with effacement of the adjacent cortical sulcus.
1b - T1 weighted post gadolinium does not show lesion enhancement.
1c - Diffusion does not show signals of restricted diffusion.
1d - FLAIR shows more irregular and heterogenous expansive lesion than previous MRI.
1e - T1 post-gadolinium shows heterogeneous lesion with impregnation by the contrast medium and contiguous abnormal meningeal enhancement.
1f and 1g - DWI and ADC maps exhibit restriction signals of the water diffusibility, related to tumor hypercellularity.
1h - T2 image shows hyperintense focal area of cystic/necrotic degeneration.
In December 2016, as the seizures persisted (focal clonic contractures of the labial commissure), the patient was hospitalized for adjustment of anticonvulsants: withdrawal of Hidantal and Phenobarbital; Clonazepam was replaced by Clobazam 10 mg 8/8 hours; and Oxcarbazepine was increased to 1,200 mg per day. A new cranial MRI showed that the left front parietal lesion had evolved to a heterogeneous form, with restriction foci (DWI) and meningeal enhancement. In January 2017 the patient presented with right hemiparesis. In February 2017 the patient underwent a biopsy. The pathological report of neovascularization and necrosis sites was consistent with glioblastoma multiforme (GBM). GBM is a highly aggressive diffuse astrocytic tumor, which is rare at a young age.
Because of rapid evolution of the patient’s symptoms, the Neurology team of HUPE requested additional molecular analysis of the tumor, and contact with the team of the Circulating Biomarkers Laboratory in UERJ was established. At biopsy, a small tumor sample was collected for molecular analyses and the proposed molecular approaches were submitted and approved by the HUPE ethical committee (number 76699717.8.0000.5259). The father of the patient signed the consent.
The Neurosurgery team did not consider surgical removal of the tumoral lesion since it spanned through eloquent areas in the left temporal and parietal cortices. Considering the rapid progression of the disease, the oncology team external to HUPE determined that treatment with the alkylating chemotherapeutic agent Temozolomide (TMZ) would not be effective for the patient. In fact, considering her decreased level of consciousness, she was referred to hospitalization at an Intensive Care Unit.
In May 2017, the patient was intubated due to the decrease in consciousness secondary to intracranial hypertension in the context of oedema and tumoral bleeding (KPS 10). The patient died on June 3, 2017. Overall survival (counting from the diagnosis date) was 4 months, although the first symptoms had emerged one year before.
Virus detection
As for the molecular approaches, DNA and RNA were extracted from the tumor sample. DNA was extracted by the Phenol-Chroroform method and the RNA by TRIzol® reagent according to the manufacturer’s instructions (InvitrogenTM).
CHIKV expression was assayed by quantitative PCR (qPCR), and the RNA was retro-transcribed into cDNA and amplified in a single step, which was accomplished by following the protocol of the PROBE One-Step RT-qPCR Kit (LGC Biotechnology) with primers provided by the Centers for Disease Control and Prevention (CDC Trioplex Real-time RT-PCR Assay Primer and Probe Set; CDC, catalog #KT0166.). The amplification was performed with the 7500 Real Time PCR System (Applied Biosystems) with a final volume of 25 μL. Also included were a negative control (no DNA) and the positive control Trioplex RT-PCR Assay Positive Control Set; CDC, catalog #KT0167.
The presence of DNA from CMV (cytomegalovirus), EBV (Epstein-Barr virus), HPV (human papillomavirus), HPyV (human polyomavirus), and VZV (of varicella-zoster virus) was screened by conventional PCR (Demmler et al., 1988; Saito et al., 1989; Manos et al., 1989; Arthur et al., 1989; Abe et al., 1998, respectively). The amplified PCR products were analyzed by 10% polyacrylamide gel electrophoresis and silver stained.
Mutation screening
Screening of mutations in the tumor suppressor gene TP53 (exons 4-10) and in the hotspots of the oncogenes NRAS/KRAS (exons 1-2) was assessed by loading onto an ABI 3730XL DNA Analyzer. Details of the sequencing reaction and the analysis can be seen in our previous publication (Leite et al., 2017).
Methylation
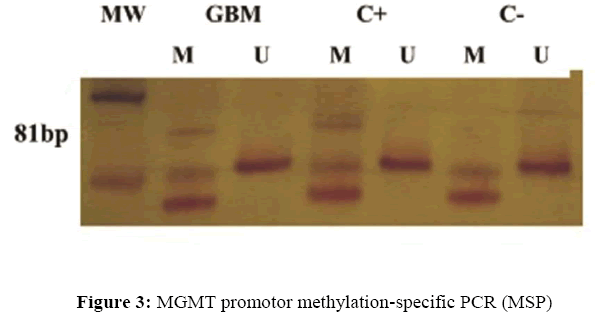
The Methylation-Specific PCR (MSP) method was applied for MGMT with the primers described previously (Esteller et al., 2000). The tumor DNA was subjected to the bisulfite conversion reaction using the EpiTect Bisulfite Kit (Qiagen, cat # 59104). The amplified PCR products were analyzed by 10% polyacrylamide gel electrophoresis and silver stained.
Results
The tumor immunohistochemical studies revealed GFAP positive in rare cells, IDH-1 positive, vimentin positive, ATRX positive, and Ki-67 positive in 35% of neoplastic cells, in addition to p53 and synaptophysin negatives. The CHIKV expression was assayed by qPCR was negative. The PCR-based test detected the presence of CMV (cytomegalovirus) alone (Figure 2). No mutations at the hotspots of NRAS/KRAS or in TP53 were detected by Sanger sequencing. The MSP-PCR test revealed hypermethylation of MGMT hypermethylation (Figure 3).
Note: The PCR products were loaded in the nondenaturing 10% polyacrylamide gels stained with silver. MW= molecular weight (Ladder 100 bp, Invitrogen Tech-Line, USA), GBM (Glioblastoma), C+ positive control, C- Negative control, GBM (Glioblastoma).
Note: The PCR products were loaded in the nondenaturing 10% polyacrylamide gels stained with silver. MW= molecular weight (Ladder 50 bp, Ludwig Biotecnologia, Brazil); GBM (Glioblastoma) - DNA methylated (M) and unmethylated (U), respectively; The DNA Methylated Universal Standard (Zymo Research, USA, cat # D5012) was used as positive control (C+); C- Negative Control.
Discussion
The CHIKV emerged in Brazil in 2014 and is transmitted through Aedes aegypti mosquito bite (Vega-Rúa et al., 2014; Nunes et al., 2015). The acute phase of CHIKV infection typically lasts from a few days to a couple of weeks. In contrast to the acute phase, the chronic phase of the disease has not been extensively investigated. Recurrent joint pain, which can last for years in some cases, is experienced by 30–40% of those infected, although this is not thought to be a result of chronic infection, as infectious virus cannot be isolated from these patients (Schwartz and Albert, 2010). CHIKV seems to infect the stromal cells of the central nervous system and, in particular, the lining of the choroid plexus. Recent outbreaks of Chikungunya have been associated with unusual severity and neurological sequelae (Rampal et al., 2007; Inglis et al., 2010; Schwartz and Albert, 2010). Additionally, CHIKV was able to infect and to activate U-87 MG cells, a glioblastoma cell line of human astrocyte origin (Abraham et al., 2013).
The CHIKV expression test was important to exclude the eventual persistence of the CHIKV expression within the tumor. However, the infection by CHIKV was closely followed by the appearance of the first symptoms and the rapid progression of the tumor, suggesting that this virus might have contributed to tumor growth by means of inflammation. The close relationship between cancer and inflammation is well known (Grivennikov et al., 2010; Hanahan and Weinberg, 2011, Hanahan and Coussens, 2012). The inflammatory microenvironment activates cells of the immune system that enhance tumor progression (Zanotto-Filho et al., 2017). CMV is a member of the Herpesviridae family, and often causes asymptomatic infections, but in some cases CMV can cause harm to human health (Slobedman and Mocarski, 1999; Brytting M et al., 1992). Evidence of the participation of CMV in gliomas has emerged recently (Cobbs et al., 2002; Cobbs, 2011; Mitchell et al., 2008; Lucas et al., 2011; Fonseca et al., 2012). An immune-based therapy has the potential to improve GBM patients’ outcome by supplementing standard treatment with chemotherapy (Schuessler et al., 2014). Unfortunately, our patient had no chance to be included in a clinical trial for CMV treatment that could have benefitted her.
The absence of mutations in TP53 was in agreement with the p53 negative immunohistochemistry result. Mutations in the oncogenes NRAS/KRAS do not represent an important prognosis or predictive biomarker in GBM; nevertheless, we assessed their DNA sequences because mutations in these oncogenes are frequent in many types of tumors, promoting tumor progression. The DNA repair protein O 6-methylguanine-DNA methyltransferase (MGMT) is important to the treatment of GBM. Active MGMT removes alkyl groups from DNA, thereby reducing the efficacy and promoting cancer cell resistance to TMZ. The test of MGMT hypermethylation (gene inactivation) was positive but this finding was useless. Unfortunately, as mentioned before, the Oncology team evaluated that due to the rapid progression of the disease (KPS 30), the TMZ therapy would be ineffective (Alonso et al., 2017).
GBM can be either pediatric or adult, or primary (developing de novo) or secondary. In the case of our patient, her age (20 years) was borderline between pediatric and adult. Considering the adults´ biomarkers, a GBM with mutation at IDH-1 (Isocitrate Dehydrogenase Gene 1) is characteristic of secondary GBM, being rare in primary GBMs (Ohgaki et al., 2016; Jiao et al., 2012). In the case of our patient, the clinical history suggests a very fast evolving tumor of astrocytic lineage ending in a secondary GBM, as the first imaging of the tumor was not suggestive of GBM, thereby making the secondary GBM hypothesis more plausible. It is possible that CHIKV and CMV had a role in the tumor’s fast progression by means of inflammation. On the other hand, mutations in TP53, which are common in the secondary adult GBMs and in the pediatric GBMs, were not detected in the tumor of our patient, making this case genetically unusual (Haynes HR, 2014).
The tumor of our patient was positive by immunohistochemistry for IDH-1 and ATRX. Alterations in IDH are found in <5% of pediatric GBMs although it is seen in adolescents with greater frequency (Haynes et al., 2014). Alterations in IDH-1 are considered an independent favorable prognosis marker, as they imply a secondary GBM; however, the altered IDH-1 did not promote a longer survival of our patient. ATRX is a critical chromatin modifier and is responsible for the lengthening of telomeres in GBM. Mutations in ATRX were found in 7% of the adult GBMs and in 14-31% of the pediatric GBMs (Jiao et al., 2012). ATRX inactivation was associated with mutations in TP53 and in IDH-1 (Jiao et al., 2012; Haynes et al., 2014). However, in our case the ATRX alteration was concordant only with the alteration in IDH-1.
Conclusion
Here we describe the case of a 20-year-old female of African descent who died due to a GBM that did not present mutations in TP53 nor in NRAS/KRAS, although MGMT hypermethylaton was positive. In contrast to most prevalent cancers, the etiology of brain tumors in adolescents is still little understood. Our patient was not subject to any known cancer risk factors, but our findings suggest the contribution of Citomegalovirus (CHIKV) and Chikungunya (CMV) for the fast tumor progression.
Disclosure
There are no conflicts of interest to declare.
About the Authors
Corresponding Author
G. Alves
Circulating Biomarkers Laboratory, Pathology Department, Medical Sciences Faculty, State University of Rio de Janeiro (UERJ), Brazil
- Email:
- galvesbrown@gmail.com
References
- Abe T, Sato M, Tamai M (1998) Correlation of varicella-zoster virus copies and final visual acuities of acute retinal necrosis syndrome. Arch Clin Exp Ophthalmol 236: 747-752. https://doi.org/10.1007/s004170050153
- Abraham R, Mudaliar P, Padmanabhan A, Sreekumar E (2013) Induction of cytopathogenicity in human glioblastoma cells by chikungunya virus. PLoS One 8:e75854. https://doi.org/10.1371/journal.pone.0075854
- Alonso D, Matallanas M, Riveros-Pérez A, Pérez-Payo M, et al. (2017) Prognostic and predictive factors in high-grade gliomas. Experience at our institution. Neurocirugia (Astur) 28: 276-283. https://doi.org/10.1016/j.neucie.2017.07.003
- Arthur RR, Dagostin S, Shah KV (1989) Detection of BK virus and JC virus in urine and brain tissue by the polymerase chain reaction. J Clin Microbiol. 27: 1174 -1179.
- Brytting M, Xu W, Wahren B, Sundqvist VA (1992) Cytomegalovirus DNA detection in sera from patients with active cytomegalovirus infections. J Clin Microbiol 30: 1937-1941.
- Cobbs CS (2011) Evolving evidence implicates cytomegalovirus as a promoter of malignant glioma pathogenesis. Herpesviridae. 2: 10. https://doi.org/10.1186/2042-4280-2-10
- Cobbs CS, Harkins L, Samanta M, Gillespie GY, et al. (2002) Human cytomegalovirus infection and expression in human malignant glioma. Cancer Research. 62: 3347-3350.
- Demmler GJ, Buffone GJ, Schimbor CM, May RA (1988). Detection of cytomegalovirus in urine from newborns by using polymerase chain reaction DNA amplification. J Infect Dis 158: 1177-1184. https://doi.org/10.1093/infdis/158.6.1177
- Esteller M, Garcia-Foncillas J, Andion E, Goodman SN, et al. (2000) Inactivation of the DNA-repair gene MGMT and the clinical response of gliomas to alkylating agents. N Engl J Med. 343: 1350-1354. https://doi.org/10.1056/nejm200011093431901
- Fonseca RF, Kawamura MT, Oliveira JA, Teixeira A, et al. (2012) The prevalence of human cytomegalovirus DNA in gliomas of Brazilian patients. Mem Inst Oswaldo Cruz 107: 953-954. https://doi.org/10.1590/s0074-02762012000700020
- Grivennikov SI, Greten FR, Karin M (2010) Immunity, inflammation, and cancer. Cell 140: 883-899. https://doi.org/10.1016/j.cell.2010.01.025
- Hanahan D, Coussens LM (2012) Accessories to the crime: functions of cells recruited to the tumor microenvironment. Cancer Cell 21: 309-322. https://doi.org/10.1016/j.ccr.2012.02.022
- Hanahan D, Weinberg RA (2011) Hallmarks of cancer: the next generation. Cell 144: 646-674. https://doi.org/10.1016/j.cell.2011.02.013
- Haynes HR, Camelo-Piragua S, Kurian KM (2014) Prognostic and predictive biomarkers in adult and pediatric gliomas: toward personalized treatment. Front Oncol 4: 47. https://doi.org/10.3389/fonc.2014.00047
- Inglis FM, Lee KM, Chiu KB, Purcell OM, et al. (2016) Neuropathogenesis of Chikungunya infection: astrogliosis and innate immune activation. J Neurovirol 22: 140-148. https://doi.org/10.1007/s13365-015-0378-3
- Instituto Nacional de Câncer. INCA. (2018). Available at [http://www.inca.gov.br/estimativa/2018/estimativa-2018.pdf].
- Jiao Y, Killela PJ, Reitman ZJ, Rasheed AB, et al. (2012) Frequent ATRX, CIC, FUBP1 and IDH1 mutations refine the classification of malignant gliomas. Oncotarget 3: 709-722. https://doi.org/10.18632/oncotarget.588
- Leite C, Delmonico L, Alves G, Gomes RJ, et al. (2017). Screening of mutations in the additional sex combs like 1, transcriptional regulator, tumor protein p53, and KRAS proto-oncogene, GTPase/NRAS proto-oncogene, GTPase genes of patients with myelodysplastic syndrome. Biomed Rep 7: 343–348. https://doi.org/10.3892/br.2017.965
- Louis DN, Perry A, Reifenberger G, von Deimling A, et al. (2016) The 2016 World Health Organization Classification of Tumors of the Central Nervous System: a summary. Acta Neuropathol 131: 803-820. https://doi.org/10.1007/s00401-016-1545-1
- Lucas KG, Bao L, Bruggeman R, Dunham K, et al. (2011) The detection of CMV pp65 and IE1 in glioblastoma multiforme. J Neurooncol. 103:231-238. https://doi.org/10.1007/s11060-010-0383-6
- Manos MM, Ting Y, Wright DK, Lewis AJ, et al. (1989) Use of polymerase chain reaction amplification for the detection of genital human papillomavirus. Cancer Cells 7: 209–214.
- Mitchell DA, Xie W, Schmittling R, Learn C, et al. (2008) Sensitive detection of human cytomegalovirus in tumors and peripheral blood of patients diagnosed with glioblastoma. Neuro Oncol 10: 10-18. https://doi.org/10.1215/15228517-2007-035
- Nunes MR, Faria NR, De Vasconcelos JM, Golding N, et al. (2015) Emergence and potential for spread of Chikungunya virus in Brazil. BMC Med 13:102. https://doi.org/10.1186/s12916-015-0348-x
- Ohgaki H, Kleihues P, Von Deimling A, Louis DN, et al. (2016) Glioblastoma, IDH mutant. In: WHO Classification of Tumours of Central Nervous System. International Agency for Research on Cancer (IARC), Lyon.
- Rampal, Sharda M, Meena H (2007) Neurological complications in Chikungunya fever. J Assoc Physicians India. 55: 765-769.
- Saito I, Servenius B, Comptn T, Fox RI (1989) Detection of Epstein-Barr virus DNA by polymerase chain reaction in blood and tissue biopsies from patients with Sjogren´s syndrome. J Exp Med 169: 2191-2198. https://doi.org/10.1084/jem.169.6.2191
- Schuessler A, Smith C, Beagley L, Boyle GM, et al. (2014) Autologous T-cell therapy for cytomegalovirus as a consolidative treatment for recurrent glioblastoma. Cancer Res. 74: 3466-3476. https://doi.org/10.1158/0008-5472.can-14-0296
- Slobedman B, Mocarski ES (1999) Quantitative analysis of latent human cytomegalovirus. J Virol. 73: 4806-4812.
- Schwartz O, Albert ML (2010) Biology and pathogenesis of chikungunya virus. Nat Rev Microbiol. 8: 491-500. https://doi.org/10.1038/nrmicro2368
- Vega-Rúa A, Zouache K, Girod R, Failloux AB, et al. (2014) High level of vector competence of Aedes aegypti and Aedes albopictus from ten American countries as a crucial factor in the spread of Chikungunya virus. J Virol 88: 6294-6306. https://doi.org/10.1128/jvi.00370-14
- Zanotto-Filho A, Gonçalves RM, Klafke K, de Souza PO, et al. (2017) Inflammatory landscape of human brain tumors reveals an NFκB dependent cytokine pathway associated with mesenchymal glioblastoma. Cancer Lett. 390: 176-187. https://doi.org/10.1016/j.canlet.2016.12.015
Keywords:
Download:
Full PDF- Share This