Population genetic structure of three species in the genus Astrocaryum G. Mey. (Arecaceae)
Received: May 25, 2017
Published: August 31, 2017
Genet.Mol.Res. 16(3): gmr16039676
DOI: 10.4238/gmr16039676
Abstract
We assessed the level and distribution of genetic diversity in three species of the economically important palm genus Astrocaryum located in Pará State, in northern Brazil. Samples were collected in three municipalities for Astrocaryum aculeatum: Belterra, Santarém, and Terra Santa; and in two municipalities for both A. murumuru: Belém and Santo Antônio do Tauá and A. paramaca: Belém and Ananindeua. Eight microsatellite loci amplified well and were used for genetic analysis. The mean number of alleles per locus for A. aculeatum, A. murumuru, and A. paramaca were 2.33, 2.38, and 2.06, respectively. Genetic diversity was similar for the three species, ranging from HE = 0.222 in A. aculeatum to HE = 0.254 in A. murumuru. Both FST and AMOVA showed that most of the genetic variation was found within populations for all three species, but high genetic differentiation among populations was found for A. aculeatum. Three loci were not in Hardy-Weinberg equilibrium, with populations of A. paramaca showing a tendency for the excess of heterozygotes (FIS = -0.144). Gene flow was high for populations of A. paramaca (Nm = 19.35). Our results suggest that the genetic diversity within populations followed the genetic differentiation among populations due to high gene flow among the population. Greater geographic distances among the three collection sites for A. aculeatum likely hampered gene flow for this species.
Introduction
The palm genus Astrocaryum G. Mey. (Arecaceae) comprises 40 species with primary distributions in tropical ecosystems of South and Central America (Kahn, 2008). The genus is comprised of many life forms, from large woody species, such as A. aculeatum G. Mey. and A. murumuru Mart., to small acaulescent palms, such as A. paramaca Mart. Because they produce fruits throughout the year, like other species in the Arecaceae, Astrocaryum spp are considered key economic species in the environments where they occur (Dransfield et al., 2008).
Although almost all species of Astrocaryum are used by humans, according to Kahn (2008) only a few have real economic potential and significant importance in local and regional trades. Of these, A. aculeatum stands out due to its abundant fruits that are widely consumed by humans in the Amazon region and to its potential to become an important crop, especially in North of Brazil (Clement et al., 2005; Kahn, 2008). Likewise, A. murumuru has been suggested by Clement et al. (2005) as an important palm for the cosmetics industry because of the high quality of its oil. In contrast with the large literature available on these two palm species, there is not much information on A. paramaca. However, because A. paramaca does not produce a central stem, its low stature could facilitate fruit harvest and could be useful in crossbreeding programs.
There are few areas with Astrocaryum plantations, and these only grow A. aculeatum. Fruit harvesting of both A. aculeatum and A. murumuru is based on the exploitation of natural populations (Clement et al., 2005; de Macêdo et al., 2015). Ferreira and Gentil (2006) suggest that difficulty in seed germination due to seed dormancy and, in the case of A. aculeatum, the inability to use vegetative propagation, contribute to a lack of interest by farmers to develop these species as crops. However, the absence of adequate management strategies for any species can have a great impact on the genetic diversity of its populations and therefore compromise its long-term survival (Booy et al., 2000).
The genetic diversity of a species can be assessed by quantifying morphological characters and biochemical or molecular markers. Of these, molecular markers offer numerous advantages over the other two types. First, they are stably expressed and detectable in all tissues regardless of growth and cell differentiation (Hamrick et al., 1992; Mondini et al., 2009). Moreover, molecular markers are not influenced by the environment, as are morphological traits (Hamrick et al., 1992). Importantly, microsatellites are especially good for studies of population genetic structure because they are highly polymorphic and allow for the final scaleresolution of population genetic parameters.
Studies of genetic diversity in palm species have been increasing in the last decades and, in most of them, the level of genetic diversity was high, with the percentage of polymorphic loci ~100%, the number of alleles per locus greater than four, and heterozygosity higher than 0.5 (e.g., Conte et al., 2003; Meerow et al., 2003; Choo et al., 2010; Gaiero et al., 2011; Gomes et al., 2011). Other population genetic structure parameters, such as Wright’s inbreeding coefficient and gene flow, have shown that, in general, genetic differentiation (FST) is low, indicating genetic variation higher within - rather than among - populations and a high gene flow among populations (Elshibli and Korpelainen, 2008; Martins-Corder et al., 2009; Shapcott et al., 2009; Namoff et al., 2011; Silva et al., 2011; Giovino et al., 2014; Nazareno and dos Reis, 2014; Oliveira et al., 2014; Moura et al., 2015). Recently, Ramos et al. (2016) found a high level of genetic diversity, mostly within populations, for adults and seedlings in populations of A. aculeatum, and along with results for spatial genetic structure indicated short-distance seed dispersal and modest levels of pollen flow. To date, there is no information regarding the level and distribution of genetic diversity for A. murumuru and A. paramaca, which is needed to evaluate the potential of them as potential crop species. Here, we present molecular genetic data to evaluate the level and distribution of genetic diversity within and among natural populations of A. aculeatum, A. murumuru, and A. paramaca.
Materials and Methods
Study species, sampling sites, and plant material
A. aculeatum is a large palm growing up to 35 m, A. murumuru is classified as a stemmed palm with a height up to 10 m, and A. paramaca is an acaulescent species (Kahn, 2008). Both A. aculeatum and A. paramaca have single stem habit, whereas A. murumuru is multi-stemmed. All three species are monoecious, perennial, produce fleshy fruits containing a single seed, and their inflorescences are composed of separate male and female flowers distributed in dyads and triads (Kahn, 2008). The mating system of A. aculeatum was classified as outcrossing (Ramos et al., 2011). To date, there is no information on the mating system of the other two species studied here. All three species have tropical distribution with common occurrence in Brazil, Guyana, and Suriname. A. murumuru and A. paramaca are also found in French Guiana, while A. aculeatum occur in Bolivia, Trinidad, and Venezuela (Kahn, 2008). In Brazil, these species are restricted to the Amazon region, in the northern part of the country (Leitman et al., 2015). Only A. aculeatum and A. murumuru have economic importance in the Amazon region, the first due to its edible fruits largely consumed in Amazonas State, and the second for its oil utilized in the national cosmetic industry (Clement et al., 2005).
We carried out sampling expeditions in three municipalities for A. aculeatum - Belterra (2°41'54''S and 54°53'18''W, 146 m above sea level, asl), Tapajós National Forest, in Santarém (2°26'22''S and 54°41'55''W, 43 m asl) and Terra Santa (2°6'16''S and 56°29'15''W, 19 m asl) - and two municipalities for both A. murumuru - Cotijuba Island (1°13'04''S and 48°32'44''W, 10 m asl) in Belém and Santo Antônio do Tauá (1°9'9''S and 48°7'45''W, 24 m asl) - and A. paramaca - Experimental fields located in Belém (1°27'18''S 48°30'9''W, 6 m asl) and Ananindeua (1°21'59''S and 48°22'20''W, 2 m asl) - all in Pará State, Brazil (Figure 1). According to Köppen’s classification, the climate in Belterra, Santarém, and Terra Santa is type Ami, Equatorial monsoon, with a minimum temperature higher than 18°C and variation of less than 5°C, and annual precipitation of 1820 mm. The climate in Belém, Santo Antônio do Tauá, and Ananindeua is classified as Afi, Equatorial rainforest (fully humid), differing from the Ami only in the annual precipitation at ~2800 mm (Kottek et al., 2006).
For the genetic diversity and population structure analyses, we collected leaves from a minimum of 20 randomly chosen adult individuals in each site. Plant material was kept on ice and transported to the Laboratory of Molecular Genetic at EMBRAPA Western Amazon, Belém, Pará, where they were stored at -20°C until DNA extraction.
DNA extraction and PCRs
For DNA extraction, leaves stored were rinsed in a 10% sodium hypochlorite solution for approximately 1 min. Total genomic DNA of each was extracted using ~100 mg leaf tissue following Costa and Oliveira (2002). Thirteen microsatellite markers developed by Ramos et al. (2012) were tested and PCR conditions were adapted for duplex and triplex reactions (Table S1). Combinations of multiplexes are shown in Table S1. PCR mix (11 μL) contained 10 ng total genomic DNA, 1.4X PCR buffer, 1.5 mM MgCl2, 0.8 μg BSA, 200 μM of each dNTP, 0.16 μM forward and M13 labeled primer (NED, FAM, and VIC), 0.32 μM reverse primer, 0.4 U Taq (5 U/μL), and milli-Q water to complete the final volume. The fragment amplifications were performed using the GeneAmp PCR System 9700 from Applied Biosystems and Mastercycler Ep gradient on Eppendorf thermal cyclers. The PCR programs were composed by two steps. The first step consisted of denaturation (68°C for 2 min and 94°C for 30 s), followed by 30 cycles [30 s at 92°C, 35 s at the primer combination-specific annealing temperature (Table S1), and 30 s at 68°C]. The second step consisted of 15 cycles (30 s at 92°C, 30 s at 53°C, and 30 s at 72°C), and a final extension at 72°C for 30 min as suggested by Ramos et al. (2012).
After PCRs had been completed, plates were prepared to contain a mix of 0.5 to 2 μL of three different multiplex products labeled with different dyes, 0.2 μL Liz 500 size standard and 10 μL Hi-Di Formamide and submitted to electrophoresis in an automated ABI 3730xl Genetic Analyzer to determine the size of the DNA fragments obtained.
Data analysis
For population genetic diversity analyses, mean number of alleles per locus (NA), mean effective number of alleles (NE), which is the number of alleles equally frequent needed to obtain the same level of homozygosity found for the real population, percentage of polymorphic loci (P%), expected heterozygosity (HE), observed heterozygosity (HO), and fixation index (F) were estimated. Estimates were obtained for each locus and means for each population and species calculated based on the average over all loci. For population genetic structure analysis, Wright F-statistics and gene flow (Nm) were computed. Deviation from Hardy-Weinberg equilibrium (HWE) was assessed for each locus, in each population, for each species using the chi-square test. Analysis of molecular variance (AMOVA) among the populations of each species was performed with two hierarchical levels (among and within populations). Principal coordinate analysis (PCoA) was performed among all individuals, for each species, after conversion of the individual-by-individual genetic distance matrix to a covariance matrix and data standardization. When applicable, a Mantel test was conducted to estimate the correlations between genetic and geographic distances. The above analyses were all conducted using the GenAlEx v.6.501 package software (Peakall and Smouse, 2006, 2012).
The assignment of genotypes to genetic clusters for each species was assessed by a Bayesian cluster analysis using the Structure 2.3.3 software (Pritchard et al., 2000). The Structure software identifies genetic clusters and assigns genotypes to those clusters without a priori information on their geographical origin. Both the admixture model and the independent allele model were run. Following a burn-in period of 100,000 runs, ten independent runs were carried out for each value of K (from 1 to 7) with 100,000 repeats. The choice of the most likely number of clusters (K) was carried out by calculating the statistics ΔK, which is based on the rate of change in the log probability of the data between successive K values, as described by Evanno et al. (2005) using Structure Harvester (Earl and Von Holdt, 2011). Among the 10 runs per K, the one with the highest maximum likelihood was used to assign individual genotypes to clusters.
Results
Results
Genetic diversity
The test for HWE suggests the occurrence of assortative mating, especially in A. paramaca populations. The chi-square test values for populations of the three species studied are shown in Table S2. Significantly deviation from HWE was found for three loci, Aac04 in populations 1 and 2 of A. aculeatum and populations 2 of A. murumuru and A. paramaca; Aac06 deviated from HWE in populations 1 and 2 of A. aculeatum and in population 1 of A.murumuru and population 2 of A. paramaca; and Aac12 deviated from HWE in population 3 of A. aculeatum and both populations of A. murumuru and A. paramaca.
Eight of the thirteen primers tested produced positive peaks allowing their fragment sizing. Considering all three species together, most primers were polymorphic. The most polymorphic primers, across all populations of the three species, were Aac04, Aac06, and Aac12 (Table S3). However, one (Aac13) was monomorphic across all species, and two were monomorphic in some species (Aac02 in A. aculeatum and A. murumuru; Aac11 in A. paramaca). The total number of alleles varied among species, ranging from 18 in populations of A. paramaca to 24 for A. aculeatum (Table S3).
The average P%, for all three species, varied from 62.5% for A. aculeatum to 75% for A. murumuru (Table 1). The mean NA ranged from 1.75 to 2.88 and the mean NE ranged from 1.32 to 1.85; all these values found for populations 1 and 2 of A. aculeatum, respectively. When the average among populations within species was considered, A. aculeatum and A. murumuru showed the highest value and similar estimates for both meanNA and NE (Table 1). Estimates of HO and HE were higher for populations 1 of A. paramaca and A. aculeatum, respectively. The mean HO and HE were higher for A. paramaca with 0.285 and A. murumuru with 0.254, respectively (Table 1). Populations 2 and 3 of A. aculeatum, 1 and 2 of A. paramaca, and population 2 of A. murumuru showed values of HO higher than the HE. The F, equivalent to FIS, varied from positive and negative among populations, being statistically different from zero for populations 1 and 3 of A. aculeatum, populations 1 and 2 of A. paramaca and population 2 of A. murumuru. When means for species are considered, only A. paramaca showed a negative F, significantly different from zero, suggesting a tendency for the excess of heterozygotes and other species that did not statistically deviate from HWE (Table 1).
| Species/Pop | N | NA | NE | P (%) | HE | HO | F (SD) |
|---|---|---|---|---|---|---|---|
| A. aculeatum | |||||||
| Belterra | 27 | 2.875 | 1.852 | 62.5 | 0.313 | 0.239 | 0.149 (0.133) |
| Santarém | 19 | 1.750 | 1.322 | 62.5 | 0.175 | 0.185 | 0.163 (0.194) |
| Terra Santa | 30 | 2.375 | 1.369 | 62.5 | 0.179 | 0.221 | -0.139 (0.083) |
| Mean | 25.333 | 2.333 | 1.514 | 62.5 | 0.222 | 0.215 | 0.058 |
| SD | 1.013 | 0.322 | 0.159 | 0 | 0.052 | 0.055 | 0.082 |
| A. murumuru | |||||||
| Cotijuba Island | 27.375 | 2.500 | 1.568 | 75 | 0.239 | 0.178 | 0.132 (0.130) |
| St. Ant. do Tauá | 30.375 | 2.250 | 1.542 | 75 | 0.269 | 0.272 | -0.060 (0.124) |
| Mean | 28.875 | 2.375 | 1.555 | 75 | 0.254 | 0.225 | 0.036 |
| SD | 0.576 | 0.287 | 0.182 | 0 | 0.062 | 0.060 | 0.089 |
| A. paramaca | |||||||
| Belém | 28.625 | 2.125 | 1.500 | 75 | 0.244 | 0.291 | -0.123 (0.061) |
| Ananindeua | 30 | 2.000 | 1.453 | 62.5 | 0.206 | 0.279 | -0.154 (0.145) |
| Mean | 29.313 | 2.063 | 1.477 | 68.75 | 0.225 | 0.285 | -0.137 |
| SD | 0.198 | 0.232 | 0.164 | 6.25 | 0.060 | 0.086 | 0.072 |
N is the number of adult individuals used for the parameters estimated; NA is the mean number of alleles per locus; NE is the effective number of alleles; HE is the expected heterozygosity; HO is the observed heterozygosity; and F is the fixation index (standard error for each population) and calculated as F = (HE - HO) / HE, representing the within-population inbreeding.
Table 1: Mean estimates of genetic diversity parameters and fixation index (F) estimated on eight loci (standard deviation) for adult individuals from natural populations of three species of Astrocaryum.
Genetic structure and gene flow
Mean values for Wright’s F-statistics provide insight into the distribution of genetic variation from individuals to among populations (Table 2). The mean reduction of heterozygosity within individuals in a subpopulation, FIS, was due to non-random mating. FIS was negative and statistically different from zero only for populations of A. paramaca, indicating an excess of heterozygotes. Our relatively low estimates of FST for all three species examined indicated that populations are not differentiated genetically.
| Statistics | Species | ||||
|---|---|---|---|---|---|
| A. aculeatum | A. murumuru | A. paramaca | |||
| FIS | 0.029 (0.115) | 0.033 | (0.123) | -0.144 (0.089) | |
| FIT | 0.107 (0.114) | 0.065 | (0.131) | -0.130 (0.090) | |
| FST | 0.089 (0.014) | 0.053 | (0.031) | 0.013 | (0.007) |
| Nm | 2.070 (0.843) | 22.641 | (14.231) | 70.527 | (28.277) |
| AMOVA | |||||
| Variation among populations | 19% | 13% | 1% | ||
| Variation within populations | 81% | 87% | 99% | ||
FIS is the inbreeding coefficient calculated within populations; FIT is the overall inbreeding coefficient related to the total sample; FST is the genetic differentiation among populations; Nm is the gene flow and estimates the proportion of migrants entering a population per generation. The results of AMOVA are percentages based on 999 permutations.
Table 2: Genetic distance matrix among 20 Poincianella pyramidalis individuals.
In fact, both Wright’s inbreeding coefficient and AMOVA for A. paramaca pointed to genetic differentiation among populations of only 1%. A. aculeatum and A. murumuru showed higher percentages of genetic differentiation among populations with AMOVA (Table 2). Estimates of Nm varied significantly among species, ranging from 2.57 for A. aculeatum to 19.35 for A. paramaca, suggesting a high level of migration constituted of pollen and seed dispersal, for A. paramaca. The low level of Nm found especially for A. aculeatum helps explain the higher degree of genetic differentiation among populations of this species (Table 2).
The Bayesian analysis of Structure 2.3.3 assigned genotypes to two populations only (K = 2), for all three species analyzed of Astrocaryum (Figure 2). In the case of A. aculeatum (Figure 2A), the number of genetic clusters found (K) was lower than the number of sampled populations, which suggests that populations may reflect a common evolutionary lineage (i.e., subdivision from a single large population) or historically high levels of Nm. The occurrence of genotype admixture is also noted for populations of A. murumuru and A. paramaca (Figure 2B and C, respectively), with A. paramaca individuals sharing around 50% of their genomes. This result can be explained by the high gene flow among populations in the genetic structure analysis for this species (Table 2).
According to the PCoA analysis performed for A. aculeatum, the first principal coordinate explained 72.41% of the genetic variance, and the second principal coordinate explained 27.59%, totalizing together 100% of the genetic variance, and population 3, Terra Santa, was more genetically divergent from the other two populations (Figure 3).
The Mantel test for three populations of A. aculeatum suggests that isolation by distance can influence the genetic divergence among the populations sampled (Figure 4).
Discussion
The number of alleles per locus found for the populations of A. aculeatum and A. murumuru species differed from that published by Ramos et al. (2012) (Table 3). For A. aculeatum, we found more monomorphic loci while for A. murumuru we found more polymorphism than the previous report, likely because our sample size was much higher. This is the first report for A. paramaca, and the number of alleles was similar to that of A. murumuru found in our study. Besides, samples from the two studies were collected in different states, and it is worth noting that some unique alleles for some loci (e.g., Aac01, Aac02, and Aac04) were found for A. aculeatum. The occurrence of these private alleles and the higher level of polymorphism were expected because our study included more sampling sites, a greater number of individuals/site, and a greater geographic extent sampled.
| Locus | A. aculeatum | A. murumuru | A. aculeatum | A. murumuru | A. paramaca | |
|---|---|---|---|---|---|---|
| Ramos et al. (2012) | Present study | |||||
| Aac01 | 2 (332-362) | 1 (338) | 1 (354) | 2 (339-354) | 2 (342-354) | |
| Aac02 | 2 (282-342) | 1 (291) | 1 (306) | 1 (306) | 2 (298-306) | |
| Aac03 | 3 (137-167) | 1 (145) | 3 (157-181) | 2 (157-162) | 2 (157-162) | |
| Aac04 | 6 (212-242) | 4 | (207-227) | 7 (207-274) | 3 (228-258) | 4 (231-263) |
| Aac06 | 8 (132-192) | 5 | (142-162) | 6 (141-196) | 5 (134-197) | 3 (143-156) |
| Aac11 | 2 (202-228) | 1 (205) | 2 (201-223) | 3 (215-252) | 1 (221) | |
| Aac12 | 5 (167-181) | 4 | (158-178) | 3 (163-195) | 5 (172-205) | 3 (179-195) |
| Aac13 | 3 (182-202) | 2 | (182-206) | 1 (204) | 1 (204) | 1 (204) |
| N | 40 | 4 | 80 | 60 | 60 | |
N is the total number of individuals sampled. Allele ranges (bp) are given in parentheses.
Table 3: Comparison of the number of alleles and allele size per locus found for species by Ramos et al. (2012) and in the present study.
The percentage of polymorphic loci found for the three Astrocaryum species was lower than that reported in other studies involving other palm species and using SSR (Conte et al., 2008; Elshibli and Korpelainen, 2008; Choo et al., 2010; Namoff et al., 2011; Ottewell et al., 2012; Giovino et al., 2014; Lanes et al., 2015; Nazareno and dos Reis, 2014). An extremely low value of polymorphism (6.17%) was reported for Livistona carinensis, an endangered palm species (Shapcott et al., 2009). However, given the wide geographic distribution of the palm species investigated in our study, we would have expected higher levels of polymorphism. Even wider geographic sampling is necessary to determine the level of polymorphism for A. aculeatum, A. murumuru, and A. paramaca.
Levels of genetic diversity (HE and HO) found for the species of Astrocaryum
presented here were also lower than most of the literature available for other palm species
using microsatellite markers, which showed a minimum estimate for HE of 0.5 (Conte et
al., 2008; Elshibli and Korpelainen, 2008; Choo et al., 2010; Namoff et al., 2011; Ottewell et al., 2012; Nazareno and dos Reis, 2014; Ramos et al., 2016). Nevertheless, similar results for HE
and HO were found for Chamaerops humilis, with a mean NA equal to 3.67 and HE and HO equal
to 0.374 and 0.366, respectively (Giovino et al., 2014). It is worth mentioning that studies
with other congeners of Astrocaryum (Oliveira et al., 2014; Ramos et al., 2012, 2016) that
used the same microsatellite set used here, excluded monomorphic loci from the estimation
of genetic diversity; this automatically increased the estimates for these parameters [given
the formula used to calculate heterozygosity 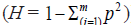 ] . Genetic diversity is dependent
on the number of alleles and the evenness of their frequencies. In fact, Conte et al. (2003) demonstrated that considering only polymorphic loci in the estimation of genetic diversity
parameters drastically affects their values. If we considered only polymorphic loci, HE and HO
estimates would represent an increase of ~60% for A. aculeatum and ~33% for A. murumuru
and A. paramaca (Table 4).
] . Genetic diversity is dependent
on the number of alleles and the evenness of their frequencies. In fact, Conte et al. (2003) demonstrated that considering only polymorphic loci in the estimation of genetic diversity
parameters drastically affects their values. If we considered only polymorphic loci, HE and HO
estimates would represent an increase of ~60% for A. aculeatum and ~33% for A. murumuru
and A. paramaca (Table 4).
| Species | NA | NE | HE | HO |
|---|---|---|---|---|
| A. aculeatum | 3.133 (0.389) | 1.823 (0.220) | 0.356 (0.061) | 0.344 (0.069) |
| A. murumuru | 2.833 (0.271) | 1.740 (0.219) | 0.338 (0.067) | 0.300 (0.066) |
| A. paramaca | 2.417 (0.229) | 1.635 (0.199) | 0.300 (0.067) | 0.380 (0.101) |
Standard deviations are shown in parentheses.
Table 4: Means for the number of alleles (NA), the effective number of alleles (NE), and expected and observed heterozygosities (HE and HO) for Astrocaryum species when only polymorphic loci are considered.
If the low levels of genetic diversity exhibited in the studied species are a result of stochastic events, such as genetic drift, population bottlenecks, or inbreeding, we would expect to find a higher proportion of homozygotes with positive values for FIS (Namoff et al., 2011). However, fixation index and Wright’s inbreeding coefficients suggested an excess of heterozygotes for A. paramaca. Other studies conducted with palm species also found a tendency for a higher proportion of heterozygotes within populations (Conte et al. (2003) ; Elshibli and Korpelainen, 2008); this could be the result of early-acting inbreeding depression mechanisms. A study with Vaccinium corymbosum (Ericaceae) found that heterozygosity increases due to the abortion of most self-fertilized seeds, a post-zygotic mechanism of high inbreeding depression (Krebs and Hancock, 1990). In a review, Husband and Schemske (1996) found general support for the theoretical prediction that, for outcrossing species, inbreeding depression tends to be expressed at early life stages. Although, there is no information on the mating system of A. murumuru and A. paramaca, close relatives, including A. mexicanum (Eguiarte et al., 1992) and A. aculeatum, are outcrossing (Ramos et al., 2011), and Oliveira NP, Oliveira MSP, Davide LC, Kalisz S (unpublished results) indicate that A. vulgare is outcrossing as well. Given the life history, floral and ecological traits shared by A. murumuru and A. paramaca with A. vulgare are likely all outcrossing.
According to our estimates of FST and AMOVA, most of the genetic variation is found within populations for all three species of Astrocaryum, following expectation for woody species (Hamrick et al., 1992). Nevertheless, genetic differentiation among A. aculetaum populations was significantly higher than for the other two species (Figure 1) likely explained by the greater distance between sample sites for A. aculetaum. Estimated gene flow for this species was much lower than for A. murumuru and A. paramaca. Similar results were found for A. aculeatum (Namoff et al., 2011) with a percentage of genetic variation within and among populations of 83.96 and 16.04, respectively. Even higher genetic differentiation among populations was found for Oenocarpus mapora (~36%), suggesting small sample sizes within populations as one possible factor to explain such variation.
Conclusion
Most of the genetic variation within the economically important palm species, Astrocaryum aculeatum, A. murumuru, and A. paramaca, is found within populations. This result is expected given their life history, floral, and ecological traits. Low levels of inbreeding occur for these species, with the proportion of heterozygotes being higher than expected by HWE in A. paramaca. Genetic differentiation among populations is strong correlated to geographic distance, with gene flow between distant populations being probably restricted by low levels of animal-mediated seed dispersal and pollen flow.
Acknowledgments
We are grateful for the assistance provided by staff of the Embrapa Eastern Amazon and University of Pittsburgh. We also would like to thank FINEP for funding the project “Pesquisa Desenvolvimento e Inovação (PD&I) em palmáceas para a produção de óleo e aproveitamento econômico de coprodutos e resíduos (Propalma, #01.10.0343.00) and CAPES for granting N.P. Oliveira a scholarship (Process #99999.001326/2014-07).
About the Authors
Corresponding Author
N.P. Oliveira
Laboratório de Citogenética, Universidade Federal de Lavras, Lavras, MG, Brasil
- Email:
- naty_poliveira@hotmail.com
References
- Booy G, Hendriks RJJ, Smulders MJM, Van Groenendael JM, et al. (2000). Genetic diversity and the survival of populations. Plant Biol. 2: 379-395. https://doi.org/10.1055/s-2000-5958
- Choo J, Ishak H, Simpson B, Mueller U, et al. (2010). Characterization of 14 microsatellite loci in a tropical palm, Attalea phalerata (Arecaceae). Am. J. Bot. 97: e105-e106. https://doi.org/10.3732/ajb.1000281
- Clement CR, Lleras Pérez E and Van Leeuwen J (2005). O potencial das palmeiras tropicais no Brasil: acertos e fracassos das últimas décadas. Agrociências 9: 67-71.
- Conte R, Nodari RO, Vencovsky R and dos Reis MS (2003). Genetic diversity and recruitment of the tropical palm, Euterpe edulis Mart., in a natural population from the Brazilian Atlantic Forest. Heredity (Edinb) 91: 401-406 https://doi.org/10.1038/sj.hdy.6800347.
- Conte R, Sedrez dos Reis M, Mantovani A and Vencovsky R (2008). Genetic structure and mating system of Euterpe edulis Mart. Populations: a comparative analysis using microsatellite and allozyme markers. J. Hered. 99: 476-482 https://doi.org/10.1093/jhered/esn055.
- Costa MR and Oliveira MSP (2002). Extração de DNA de açaizeiro a partir de folhas. Documentos. Embrapa Amazônia Oriental, Belém, n.127, 1-22.
- Dransfield J, Uhl NW, Asmussen CB, Baker WJ, et al. (2008). Genera Palmarum: the evolution and classification of palms. 2nd edn. Kew Publishing, Royal Botanical Garden, Londres, 732.
- De Macêdo JLV, Ramos SLF, Lopes MTG, da Costa JR, et al. (2015) Tucumã-do-amazonas. In: Palmeiras nativas do Brasil(Lopes R, Oliveira MSP, Cavallari MM, Barbieri RL, et al., eds.).
- Earl DA and Von Holdt BM (2011). STRUCTURE HARVESTER: A website and program for visualizing STRUCTURE output and implementing the Evanno method. Conserv. Genet. Resour. 4: 359-361. https://doi.org/10.1007/s12686-011-9548-7
- Eguiarte LE, Perez-Nasser N and Piñero D (1992). Genetic structure, outcrossing rate and heterosis in Astrocaryum mexicanum (tropical palm): Implications for evolution and conservation. Heredity https://doi.org/10.1038/hdy.1992.119.
- Elshibli S and Korpelainen H (2008). Microsatellite markers reveal high genetic diversity in date palm (Phoenix dactylifera L.) germplasm from Sudan. Genetica 134: 251-260 https://doi.org/10.1007/s10709-007-9232-8.
- Evanno G, Regnaut S and Goudet J (2005). Detecting the number of clusters of individuals using the software STRUCTURE: a simulation study. Mol. Ecol. 14: 2611-2620. https://doi.org/10.1111/j.1365-294X.2005.02553.x
- Ferreira SAN and Gentil DFO (2006). Extração, embebição e germinação de sementes de tucumã (Astrocaryum aculeatum). Acta Amazon. 36: 141-146. https://doi.org/10.1590/S0044-59672006000200002
- Gaiero P, Mazzella C, Agostini G, Bertolazzi S, et al. (2011). Genetic diversity among endangered Uruguayan populations of Butia Becc. species based on ISSR. Plant Syst. Evol. https://doi.org/10.1007/s00606-010-0412-0.
- Giovino A, Scibetta S, Saia S and Guarino C (2014). Genetic and morphologic diversity of European fan palm (Chamaerops humilis L.) populations from different environments from Sicily. Bot. J. Linn. Soc. 176: 66-81. https://doi.org/10.1111/boj.12195
- Gomes LRP, Lopes MTG, Bentes JLS, Barros WS, et al. (2011). Genetic diversity in natural populations of Buriti (Mauritia flexuosa L.f.). Crop Breed. Appl. Biotechnol. https://doi.org/10.1590/S1984-70332011000300003.
- Hamrick JL, Godt MJW and Sherman-Broyles SL (1992). Factors influencing levels of genetic diversity in woody plant species. New For. 6: 95-124. https://doi.org/10.1007/BF00120641
- Husband BC and Schemske DW (1996). Evolution of the magnitude and timing of inbreeding depression in plants.Evolution 50: 54-70. https://doi.org/10.1111/j.1558-5646.1996.tb04472.x
- Kahn F (2008). The genus Astrocaryum. Rev. Peru. Biol. 15: 31-48.
- Kottek M, Grieser J, Beck C, Rudolf B, et al. (2006). World-map of Köppen-Gieger climate classification updated. Meteorol. Z. 15: 259-263. https://doi.org/10.1127/0941-2948/2006/0130
- Krebs SL and Hancock JF (1990). Early-acting inbreeding depression and reproductive success in the highbush blueberry,Vaccinium corymbosum L. Theor. Appl. Genet. 79: 825-832. https://doi.org/10.1007/BF00224252
- Lanes ECM, Motoike SY, Kuki KN, Nick C, et al. (2015). Molecular characterization and population structure of the macaw palm, Acrocomia aculeata (Arecaceae), ex situ germplasm collection using microsatellites markers. J. Hered. 106: 102-112 https://doi.org/10.1093/jhered/esu073.
- Leitman P, Soares K, Henderson A, Noblick L, et al. (2015). Arecaceae in Lista de Espécies da Flora do Brasil. Jardim Botânico do Rio de Janeiro. Available at [http://floradobrasil.jbrj.gov.br/jabot/floradobrasil/FB22080].
- Martins-Corder MP, Fialho LEB, Zambiazi DC and Konzen ER (2009). Análise da diversidade genética de populações de palmiteiro (Euterpe edulis MARTIUS) através de marcadores isoenzimáticos. Rev. Ceres 56: 204-213.
- Meerow AW, Wisser RJ, Brown JS, Kuhn DN, et al. (2003). Analysis of genetic diversity and population structure within Florida coconut (Cocos nucifera L.) germplasm using microsatellite DNA, with special emphasis on the Fiji Dwarf cultivar. Theor. Appl. Genet. 106: 715-726 https://doi.org/10.1007/s00122-002-1121-z.
- Mondini L, Noorani A and Pagnotta MA (2009). Assessing plant genetic diversity by molecular tools. Diversity (Basel) 1: 19-35. https://doi.org/10.3390/d1010019
- Moura EF, Oliveira MSP, Silva DT and Pontes LCG (2015). Genetic diversity and structure of Oenocarpus mapora germplasm conserved at eastern Amazon. Rev. Bras. Frutic. 37: 984-992.
- Namoff S, Veloz A, Jiménez F, Rodríguez-Peña RA, et al. (2011). Sweet drinks are made of this: conservation genetics of an endemic palm species from the Dominican Republic. J. Hered. 102: 1-10 https://doi.org/10.1093/jhered/esq118.
- Nazareno AG and dos Reis MS (2014). At risk of population decline? An ecological and genetic approach to the threatened palm species Butia eriospatha (Arecaceae) of Southern Brazil. J. Hered. 105: 120-129 https://doi.org/10.1093/jhered/est065.
- Oliveira LDS, Ramos SLF, Lopes MTG, Dequigiovanni G, et al. (2014). Genetic diversity and structure of Astrocaryum jauari (Mart.) palm in two Amazon river basins. Crop Breed. Appl. Biotechnol. https://doi.org/10.1590/1984- 70332014v14n3a25.
- Ottewell K, Grey E, Castillo F and Karubian J (2012). The pollen dispersal kernel and mating system of an insectpollinated tropical palm, Oenocarpus bataua. Heredity (Edinb) 109: 332-339 https://doi.org/10.1038/hdy.2012.40.
- Peakall R and Smouse PE (2006). GenAlEx 6: genetic analysis in Excel. Population genetic software for teaching and research. Mol. Ecol. 6: 288-295. https://doi.org/10.1111/j.1471-8286.2005.01155.x
- Peakall R and Smouse PE (2012). GenAlEx 6.5: genetic analysis in Excel. Population genetic software for teaching and research - an update. Bioinformatics 28: 2537-2539. https://doi.org/10.1093/bioinformatics/bts460
- Pritchard JK, Stephens M and Donnelly P (2000). Inference of population structure using multilocus genotype data.Genetics 155: 945-959.
- Ramos SLF, Lopes MTG, Lopes R, Cunha RNV, et al. (2011). Determination of the mating system of Tucumã palm using microsatellite markers. Crop Breed. Appl. Biotechnol. https://doi.org/10.1590/S1984-70332011000200011.
- Ramos SLF, de Macêdo JL, Lopes MTG, Batista JS, et al. (2012). Microsatellite loci for tucumã of Amazonas (Astrocaryum aculeatum) and amplification in other Arecaceae. Am. J. Bot. 99: e508-e510 https://doi.org/10.3732/ajb.1100607.
- Ramos SLF, Dequigiovanni G, Sebbenn AM, Lopes MTG, et al. (2016). Spatial genetic structure, genetic diversity and pollen dispersal in a harvested population of Astrocaryum aculeatum in the Brazilian Amazon. BMC Genet. 17: 63 https://doi.org/10.1186/s12863-016-0371-8.
- Shapcott A, Dowe JL and Ford H (2009). Low genetic diversity and recovery implications of the vulnerable Bankoualé palm Livistona carinensis (Arecaceae), from North-eastern Africa and the Southern Arabian Peninsula. Conserv. Genet. https://doi.org/10.1007/s10592-008-9582-5.
- Silva MS, Vieira FA and Carvalho D (2011). Diversity and genetic structure in natural populations of Geonoma schottiana Mart. (Arecaceae): Implications for conservation. Cerne https://doi.org/10.1590/S0104-77602011000200006.
Keywords:
Download:
Full PDF- Share This



