Morphological, cytogenetic, and molecular characterization of Arachis kuhlmannii Krapov. & W.C. Greg. (Leguminosae)
Received: May 23, 2017
Accepted: August 24, 2017
Published: September 21, 2017
Genet.Mol.Res. 16(3): gmr16039733
DOI: 10.4238/gmr16039733
Abstract
Arachis kuhlmannii occurs in Mato Grosso and Mato Grosso do Sul States, Brazil. Its area of occurrence partially overlaps with that of other species in the Arachis section. Because of their morphological similarities, these species are often mistaken one for another. This study aimed the correct classification of available accessions as Arachis kuhlmannii, or other species, and the characterization of similarities among accessions and Arachis hypogaea by morphological, cytogenetic, and molecular marker analyses. Thirty-eight accessions were used. Principal component analysis was used for morphological characterization, root tips for mitotic metaphase analysis, and RAPD markers for molecular characterization. Cluster analysis discriminated accessions with the A genome from the B genome. Cluster analysis based on molecular markers discriminated natural populations in a manner that correlated with geographical areas of the collection. Arachis cardenasii and A. hypogaea were isolated from other A-genome accessions. Cytogenetic analyses confirmed the existence of diagnostic characteristics that distinguish species with the A genome from those with the B genome. Results suggest the need for a taxonomic review of some species in the Arachis section, as we could not discriminate as distinct species all of the accessions identified as A. kuhlmannii, A. helodes, and A. simpsonii by using morphological, molecular, and cytogenetic markers.
Introduction
The perennial species Arachis kuhlmannii Krapov. & W.C. Greg produces hybrids when crossed with the cultivated peanut A. hypogaea L. The species naturally occurs in the States of Mato Grosso and Mato Grosso do Sul, in the West Central region of Brazil (Krapovickas and Gregory, 1994). It represents one of the few wild species found on the entire border of the Brazilian Pantanal ecosystem, and even in the ecologically peculiar conditions of central Pantanal (Pantanal of Nhecolândia), which is characterized by a mosaic of floodable grasslands, permanent or temporary ponds, some with slightly alkaline water, savannas, and forests (Pott et al., 2011).
This distribution area partly overlaps with those of other species of the taxonomic section Arachis , such as A. cardenasii Krapov. & W.C. Greg., A. helodes Mart. ex Krapov. & Rigoni, and A. simpsonii Krapov. & W.C. Greg., all of which, in the past, have been mistaken for A. kuhlmannii. In fact, the difficulty in circumscribing the different species has led Antonio Krapovickas and Walton C. Gregory to wrongly group accessions they thought belonged to a single species (Krapovickas and Gregory, 1994). Additionally, the same authors classified other accessions with similar morphological characteristics as related materials (Arachis aff. kuhlmannii).
A previous study has demonstrated that A. kuhlmannii accessions V 8889 and V 7639 display resistance to fungal diseases including late and early leaf spot and rust (Fávero et al., 2009) and insects such as thrips and red-necked peanutworm (Michelotto et al., 2013). Therefore, understanding the complex set of plants currently grouped as A. kuhlmannii and related accessions would shed light on the taxonomy of this species, and provide potentially useful information for breeding programs.
In this study, we use morphological, cytogenetic, and molecular markers to evaluate accessions at least preliminarily allocated to the species A. kuhlmannii or in close relationship to it, to accommodate each of these accessions in the appropriate taxa. Moreover, we investigated the proximity between these accessions, A. hypogaea and other diploid species of the same taxonomic section.
Materials and Methods
Germplasm accessions
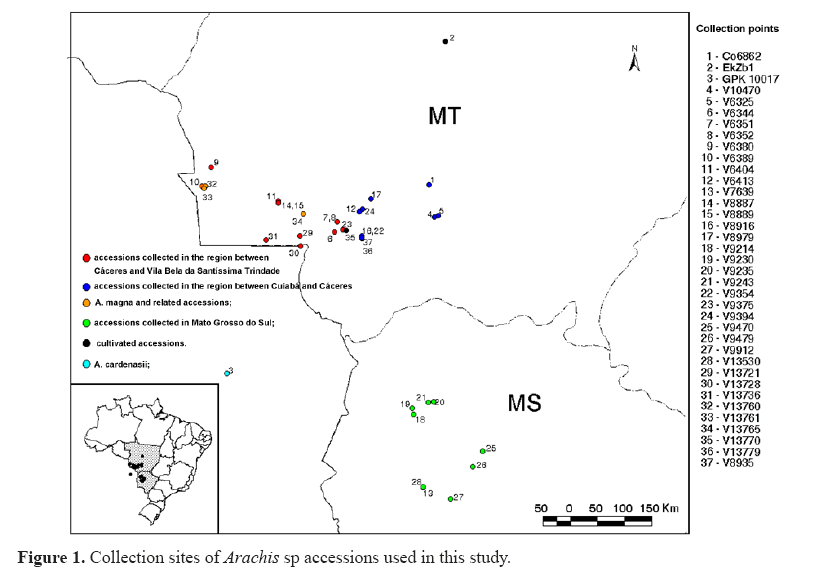
Accessions used in this study were obtained from the Wild Arachis Germplasm Bank at EMBRAPA Genetic Resources and Biotechnology, Brasília, DF, Brazil, and are listed in Table 1. Figure 1 shows the sites where each accession was collected. The study was carried out at Embrapa Genetic Resources and Biotechnology, under greenhouse conditions. Experiments were conducted with 37 germplasm accessions in three randomized complete block designs. Regarding accession V 8935, only one replication was performed. Thus, in total, 38 accessions were used in our experiments.
| Collector numbers | Brazilian code | Origin | Municipality | Identification before this study | A chromosome | Identification supported by this study |
| EkZb 1 | 031879 | MT | L. do Rio Verde | A. kuhlmannii | + | A. kuhlmannii |
| VSGr 6344 | 012599 | MT | Cáceres | A. kuhlmannii * | + | A. kuhlmannii |
| VSGr 6352 | 012611 | MT | Cáceres | A. kuhlmannii * | + | A. kuhlmannii |
| VSGr 6404 | 012653 | MT | Porto Esperidião | A. kuhlmannii * | + | A. kuhlmannii |
| VRGeSv 7639 | 017515 | MS | Miranda | A. kuhlmannii * | + | A. kuhlmannii |
| VKSSv 8887 | 020206 | MT | Porto Esperidião | A. kuhlmannii * | + | A. kuhlmannii |
| VKSSv 8889 | 020222 | MT | Porto Esperidião | A. kuhlmannii * | + | A. kuhlmannii |
| VRSSv 8916 | 020257 | MT | Cáceres | A. kuhlmannii * | + | A. kuhlmannii |
| VKSSv 8935 | 020265 | MT | Cáceres | A. kuhlmannii * | + | A. kuhlmannii |
| VPoBi 9214 | 022535 | MS | Corumbá | A. kuhlmannii * | + | A. kuhlmannii |
| VPoBi 9230 | 022543 | MS | Corumbá | A. kuhlmannii * | + | A. kuhlmannii |
| VPoBi 9235 | 022551 | MS | Corumbá | A. kuhlmannii * | + | A. kuhlmannii |
| VPoBi 9243 | 022560 | MS | Corumbá | A. kuhlmannii * | + | A. kuhlmannii |
| VPoBi 9354 | 020257 | MT | Cáceres | A. kuhlmannii * | + | A. kuhlmannii |
| VPoBi 9375 | 022594 | MT | Cáceres | A. kuhlmannii * | + | A. kuhlmannii |
| VPoBi 9470 | 022578 | MS | Aquidauana | A. kuhlmannii * | + | A. kuhlmannii |
| VPoBi 9479 | 022586 | MS | Aquidauana | A. kuhlmannii * | + | A. kuhlmannii |
| VSW 9912 | 022900 | MS | Aquidauana | A. kuhlmannii * | + | A. kuhlmannii |
| VPzRcSgSv 13530 | 032654 | MS | Miranda | A. kuhlmannii | + | A. kuhlmannii |
| V13770 | 012564 | MT | Cáceres | A. kuhlmannii * | + | A. kuhlmannii |
| VKSSv 13779 | 033871 | MT | Cáceres | A. kuhlmannii * | + | A. kuhlmannii |
| VSGr 6351 | 012602 | MT | Cáceres | A. aff. kuhlmannii * | + | A. kuhlmannii |
| VSGr 6380 | 012645 | MT | V.B.S.Trindade | A. aff. kuhlmannii * | + | A. kuhlmannii |
| VSGr 6413 | 012688 | MT | Cáceres | A. aff. kuhlmannii * | + | A. kuhlmannii |
| VKSSv 8979 | 020354 | MT | Cáceres | A. aff. kuhlmannii * | + | A. kuhlmannii |
| VPoBi 9394 | 022624 | MT | Cáceres | A. aff. kuhlmannii * | + | A. kuhlmannii |
| VSGr 6389 | 012696 | MT | V.B.S.Trindade | A. gregoryi | - | A. gregoryi |
| V13765 (=VSGr6408b) | 034011 | MT | Cáceres | A. magna | - | A. magna |
| CoSzSv 6862 | 018619 | MT | Cuiabá | A. helodes * | + | A. helodes |
| VSGr 6325 | 012505 | MT | S. Antonio do Leverger | A. helodes * | + | A. helodes |
| VPoJSv 10470 | 024945 | MT | N.S. Livramento | A. helodes * | + | A. helodes |
| GKP 10017 | 013404 | BOL | Roboré | A. cardenasii * | + | A. cardenasii |
| VKSSv 13721 | 033723 | MT | Porto Esperidião | A. aff. cardenasii ** | + | A. kuhlmannii |
| VKSSv 13736 | 033766 | MT | Porto Esperidião | A. aff. cardenasii ** | + | A. kuhlmannii |
| VPzSgRcSv 13728 | 033740 | BOL | San Matias | A. simpsonii * | + | A. simpsonii |
| VSPmSv 13760 | 033839 | MT | V.B.S.Trindade | A. magna | - | A. magna |
| VSPmSv 13761 | 033847 | MT | V.B.S.Trindade | A. aff. magna | - | A. magna |
| Cultivar Tatu | 011606 | SP | Campinas | A. hypogaea | + | A. hypogaea |
*Identifications by Krapovickas and Gregory, 1994. Collectors: Bi = L.B. Bianchetti; Co = L. Coradin; Ek = E. Kornelius; G = W.C. Gregory; Ge = M.A.N. Gerin; Gr = A. Gripp; J = L. Jank; K = A. Krapovickas; P = J.R. Pietrarelli; Pm = R.N. Pittman; Po = A. Pott; Pz = E.A. Pizarro; Rc = R.C. Oliveira; S = C.E. Simpson; Sg = A.K. Singh; Sv = G.P. Silva; Sz = R. Schultze-Kraft; W = W.L. Werneck; V = J.F.M. Valls; Zb = J.L. Zoby.
Table 1: Collector numbers, Brazilian code, the state of origin and municipality, identification of the name of species before and supported by this study, and the presence of the A chromosome in Arachis sp accessions.
Morphological characterization
Approximately 130 days after seed germination, morphological characteristics were evaluated based on descriptors traditionally used for wild species Arachis (IBPGR, 1990; Monçato, 1995). Evaluated descriptors included: epicotyl length, stem thickness, height of the main axis, lateral branch length, internode length, petiole and rachis length, basal and distal leaflet length and width, leaflet shape (obovate or lanceolate), length and width of adnate and free part of stipule, number of veins in stipule, presence of trichomes and bristles on pegs, stipule margin, stipule center, leaf margin, epiphyll, hypophyll, midvein of hypophyll, petiole, rachis, internode, hypanthium and wing width and height, lower and upper calyx lip length, color of the wings and calyx, presence of anthocyanin in hypanthium, pulvinus, peg, branch, calyx, length and width of 10 fruit segments and 10 seeds per plant (whenever possible), reticulation of fruit segments, presence/absence of isthmus between fruit segments, average seed yield per plant, annual, biennial, or perennial life cycle.
Two leaves per plant were measured: the fourth oldest leaf (bottom-up) of the main axis and the fourth youngest leaf (from the apex to the base) of the lateral branch. Colors were determined with the range of Kornerup and Wanscher (1979).
From the 78 descriptors evaluated, only 14 remained as the most important in the discrimination of accessions: main axis height (cm), annual performance (+ or -), biennial (+ or -) or perennial plants (+ or -), internode length in the first lateral branches (mm), length of branches (cm), fruit-segment length (mm), seed length (mm), epicotyl length (cm), width of fruit segments (mm), seed width (mm), presence of isthmus (+ or -), average production of seed per plant, and reticulation of fruit segment (+ or -). Hussaini et al. (1977), in a study of Eleusine coracana (L.) Gaertn., noted that 11 of 29 initially defined characteristics should be discarded. Eliminated characteristics were mostly vegetative and highly affected by environmental factors, whereas reproductive components associated with panicle traits were less affected.
Qualitative characteristics provide important evidence for differentiation among accessions. Thus, we transformed quantitative data into a binary system during cluster analysis, so that we could use both quantitative and qualitative characteristics together. To this end, we obtained the average values for each descriptor and then assigned 0 to values below the average and 1 to those above. This methodology reduced sensitivity in the discrimination of data but provided the simplest tool to group characteristics that had different evaluation metrics. For the clustering technique, we used the unweighted pair group method with arithmetic averages (UPGMA). Simple matching was used as the similarity coefficient because the absence of a descriptor is also considered a similarity. The product-moment correlation coefficient (r) was used to test the adequacy of the group related to the matrix of data. When the correlation coefficient (r) is above 0.7, the cophenetic matrix and the similarity matrix are close and provide confidence in the clusters presented by the dendrogram. Cluster analysis was performed using the NTSYS-pc 2.0 computer program (Numerical Taxonomy System of Multivariate Statistical Programs).
Cytogenetic characterization
Cytogenetic characterization focused on the presence or absence of the A chromosome pair. In species showing this pair, it appears, during metaphase, as the smallest pair of chromosomes in the cell, usually just a little above half the length of the longest pair, and stains less intensively than other chromosomes during prophase (Fernández and Krapovickas, 1994).
For cytogenetic analysis, roots ranging between 0.5-1 cm in length were collected at 9:30 am and pre-treated in mono-bromonaphthalene saturated solution for 2 h, at room temperature. The material was fixed in Carnoy solution (absolute ethanol:glacial acetic acid, 3:1, v/v) for 24 h, and stored in 70% ethanol at 4°C. For analysis of mitotic chromosomes, root tips were stained following Feulgen’s method, with hydrolysis of 1 N HCl solution, at 60°C for 10 min, and then the radicular meristems were squashed in a drop of 2% propionic carmine for intensifying the staining. Images of mitotic chromosomes were captured at Zeiss microscope following KS300 program or photographed with Ilford Pan F 50 film.
Molecular characterization
All 112 individual leaves were analyzed by RAPD markers. Leaf DNA was extracted according to the protocol described by Lodhi et al. (1994), with modifications. After extraction, 200 μL TE + RNase was added to each tube. DNA was quantified on 0.8% agarose gels. Final samples had concentrations of 2.5 ng/μL. The PCR was based on the protocol of Williams et al. (1990), with modifications by Ferreira and Grattapaglia (1996). We used 1.5% agarose gels to separate fragments. Primer screening was performed with individuals of the V 13779, V 13760, and IAC Tatu-ST accessions, which were likely the most different materials as they were supposed to have the A, B, and AB genomes, respectively. Fifty-eight primers from kits A, B, W, X, and Y by OPERON were tested. Forty primers were selected based on the number and quality of polymorphic bands (strong and without drag) for analysis of molecular markers.
During molecular characterization, we observed that one individual had a different band pattern from the other two individuals in the same accession. Thus, we separated the analyses of this plant, V 13770 belonging to block A, and added 1 OTU to the study that now totaled 39.
A data matrix was created with 113 individuals and 90 polymorphic markers. We used the Jaccard similarity coefficient because the absence of a descriptor could not be recorded as a similarity. For the clustering technique, we used UPGMA. The coefficient of productmoment correlation (r) was used to test the adequacy of the group related to the matrix data. Cluster analysis was performed using the NTSYS 2.0 computer program.
Results and Discussion
Morphological characterization
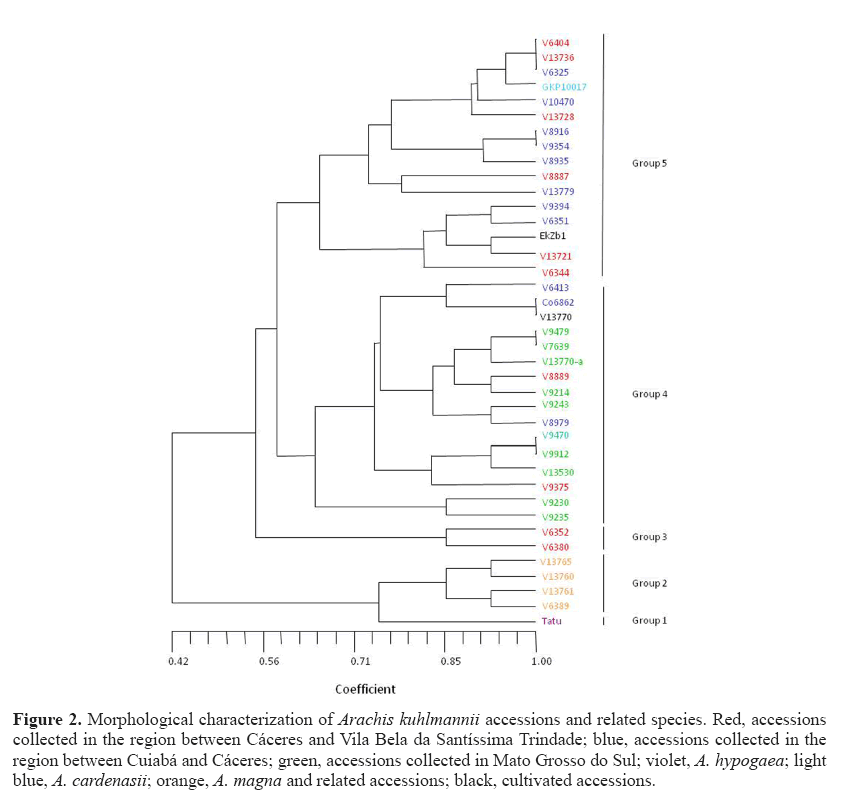
The dendrogram (Figure 2) had a cophenetic correlation coefficient (r) of 0.77, indicating a reliable grouping. Cultivated peanuts, A. hypogaea ‘Tatu’, were isolated from other accessions in this study and formed group 1, with a similarity coefficient of 0.74 concerning the nearest group (group 2), which include two subgroups identified as A. magna Krapov., W.C. Greg. & C.E. Simpson and A. gregoryi C.E. Simpson, Krapov. & Valls. The proximity between the cultivated peanut A. magna and A. gregoryi accessions can be deduced from the fact that they are all annual, and their main axis and epicotyl are higher than those of other materials. A descriptor that separates groups 1 and 2 is the absence of isthmus in the fruit of A. hypogaea, an important characteristic in the differentiation from other accessions gained in the process of domestication (Valls et al., 1995). These two groups possessed a coefficient of similarity with all other accessions observed in the dendrogram of 0.42, which shows that groups 1 and 2 are morphologically very different from all others. Two subgroups were formed within group 2, one including accessions V 13760 and V 13765 and, the other, accessions V 6389 and V 13761. This subgrouping resulted from observations of branch and internode length, both of which were larger in the second subgroup.
Figure 2: Morphological characterization of Arachis kuhlmannii accessions and related species. Red, accessions collected in the region between Cáceres and Vila Bela da Santíssima Trindade; blue, accessions collected in the region between Cuiabá and Cáceres; green, accessions collected in Mato Grosso do Sul; violet, A. hypogaea; light blue, A. cardenasii; orange, A. magna and related accessions; black, cultivated accessions.
Group 3 placed together two accessions recognized as A. kuhlmannii (V 6352) and A. aff. kuhlmannii (V 6380). Initially, we would have grouped them with other accessions of A. kuhlmannii or related material, but descriptors such as plant height and length of epicotyl, when jointly observed, isolated these materials in a group with a similarity coefficient of 0.55 in comparison to the others in the dendrogram.
Group 4 joined fifteen accessions identified as A. kuhlmannii or A. aff. kuhlmannii and Co 6862, this tentatively identified as A. helodes . Most of these materials were collected in Mato Grosso do Sul, Brazil, except Co 6862. Despite some apparently significant differences, some accessions collected in Mato Grosso (red or blue) also fell into group 4 because of their large fruit and seed size, longer branches and high seed productivity. Gregory et al. (1980) pointed to limitations in herbarium specimens deposited in one of the largest herbaria in the world, and to a disproportionate emphasis placed on vegetative parts that have strong similarities among distantly related taxa. These authors state that insufficient attention has been given to the collection and preparation of the reproductive and underground parts of voucher specimens, which are more stable, distinct and useful in the differentiation of taxonomic entities of the genus Arachis . In fact, in the current study, fruit segment and seed size, and fruit segment reticulation represented characteristics of crucial importance for the discrimination of accessions of the B genome.
Group 5 displayed a 0.58 coefficient of similarity related to group 4. Group 5 included genotypes collected in Mato Grosso State and A. cardenasii , collected in Bolivia. These accessions were grouped because of similar characteristics, such as perennial behavior, low seed yield per plant, as well as small fruit segments and seeds. A group of three species previously known to be distinct from A. kuhlmannii and a priori expected to be isolated from the other accessions included A. cardenasii (GKP 10017), A. helodes (V 6325 and V 10470), and A. simpsonii (V 13728). These species fell into group 5.
We had expected greater morphological distinction among species, especially those possessing the A genome, whose accessions were in groups 3, 4, and 5. However, morphological characterization by clustering analysis explained only three major groupings: 1) A. hypogaea ‘Tatu’; 2) containing accessions identified as A. magna or related to it; and 3) containing A-genome accessions belonging to the species A. kuhlmannii or A. aff. kuhlmannii, A. simpsonii , A. cardenasii , and A. helodes .
Monçato (1995) retained 23 descriptors with a better discriminating capacity of 52. The author separated A. pintoi Krapov. & W.C. Greg. and A. repens Handro by principal component analysis, keeping intermediary accessions as a single independent taxonomic classification between the two species. The most important parameters were stolon thickness, standard length, bristles on the rachis and petiole, fruit length and weight, and seed length, width and weight. Morphologically, A. helodes individuals have small, hairless leaflets (Krapovickas and Gregory, 1994). We observed small leaflet size in A. helodes , but the difference was not large enough to isolate the species. In fact, in Arachis species classification, leaflet length and width are usually not considered independently, but in the ratio length/width, which attaches greater importance to leaflet form than size. Hairs were present on the hypophyll of A. helodes . A. simpsonii individuals have small seeds and epiphyll hair (Krapovickas and Gregory, 1994). A. kuhlmanni accessions often have small seeds, sometimes only slightly larger than those seen in A. simpsonii . The average seed length of A. kuhlmanni V 13779, collected near the site of a collection of the type specimen KG 30034, was 10.06 mm, whereas A. simpsonii seeds were 9.02 mm long. In the monograph dichotomous key (Krapovickas and Gregory, 1994), fruit segment size ranges in length between 7 and 11 mm, and in width between 4 and 6 mm, to account for the variability in A. simpsonii . The A. kuhlmanni accessions V 13779 and V 6404 would fit in this category. In dichotomous keys, branch overlapping for characteristics such as leaflet shape, seed size, and others can make species identification more difficult. Some accessions of A. kuhlmannii, such as V 6380, exhibited short epiphyll hairiness. Some of the main features of A. cardenasii include smaller fruits than those of A. kuhlmannii and well-rounded leaflets. The many overlapping traits described here pose an obstacle to the morphological identification of Arachis species. Thus, other markers must come into play to understand such a complex group.
Cytogenetic characterization
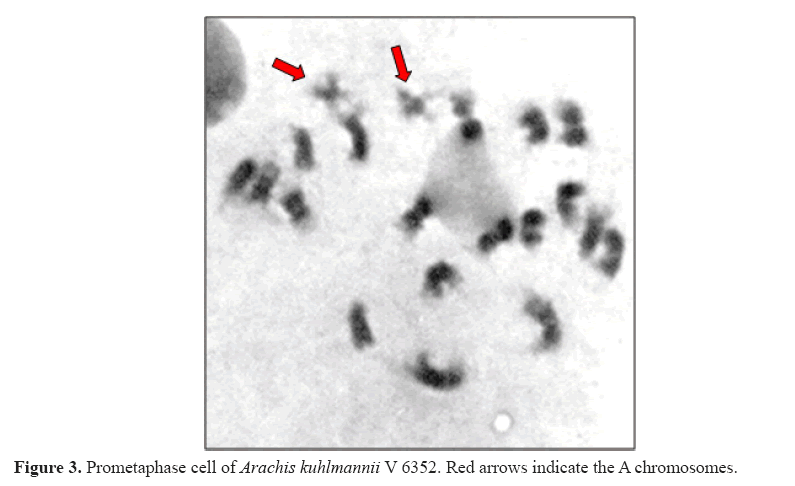
Through cytogenetic analysis, it was possible to confirm the somatic chromosome number (2n = 20) and the presence of the small A pair of chromosomes, due to its typically heteropycnotic and differential constriction at prometaphase, as observed for A. kuhlmannii V 6352 (Figure 3). At the accession of A. gregoryi V 6389, as well as accessions of A. magna V 13765, V 13761, and V 13760, the small A pair was absent. As expected, these both species were grouped, but isolated from the other analyzed species, which have A pair of chromosomes and are associated with the A genome of A. hypogaea.
Except for A. hypogaea, all accessions displayed 20 chromosomes. Table 1 lists accessions and A pair status. Tallury et al. (2005) also grouped A. kuhlmannii with A-genome accessions, both by cytogenetic (meiosis) and molecular (AFLP) characterization. The same was observed by Gimenes et al. (2007) using microsatellite markers. In a detailed cytological study, using fluorescent in situ hybridization, but including a single A. kuhlmannii accession (V 9235, also represented here), Robledo et al. (2009) classified the A pair diploid species of the Arachis section into three groups (La Plata River Basin, Chiquitano, and Pantanal). A. kuhlmannii was considered a member of the Pantanal group, alongside with A. helodes and A. simpsonii , while A. cardenasii was maintained in the distinct Chiquitano group.
Molecular analysis
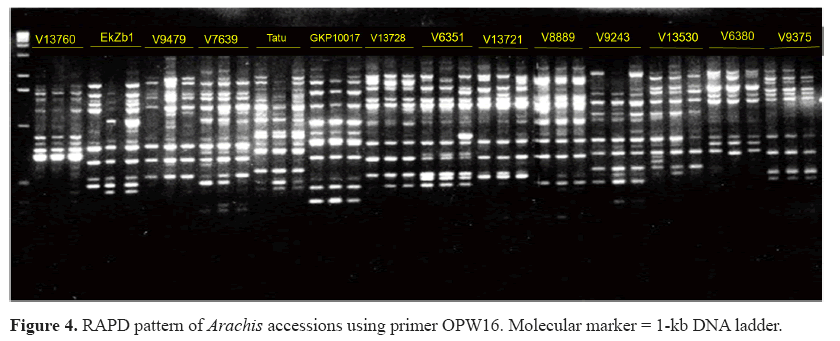
Among 58 primers tested, 14 were analyzed and 90 polymorphic markers were obtained. There was an average of seven polymorphic bands per primer. The minimum number of bands per primer was two for primers Y03 and Y05, and the maximum was 13 for primer W16. Figure 4 displays molecular profiles obtained with primer W16.
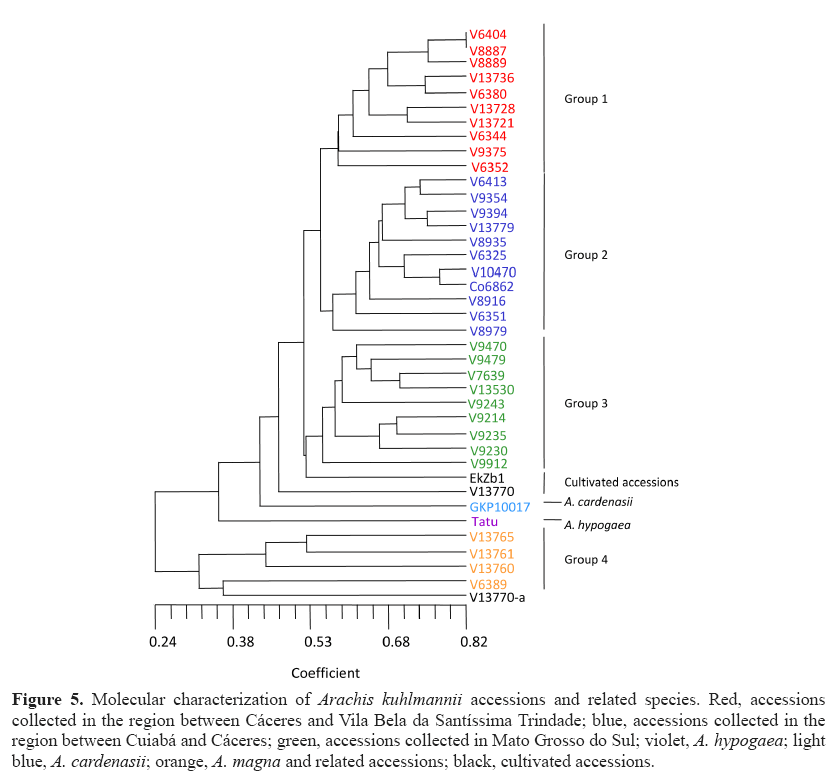
Figure 5 displays a dendrogram forming several groups. Group 1 included accessions collected in Cáceres and Vila Bela da Santíssima Trindade. Among these accessions, some were previously identified as A. kuhlmannii, A. aff. kuhlmannii, and A. simpsonii (V 13728). Groups 1 and 2 had a genetic similarity of 0.55. Nevertheless, within group 1, similarities were as low as 0.59 between V 9375 and the other materials. Thus, we believe that accessions in groups 1 and 2 may belong to the same species. Bechara et al. (2010) using ITS and 5.8S markers also noted the similarity among accessions of A. kuhlmannii and accessions of A. simpsonii and A. helodes . Group 2 included accessions collected in the region of Cáceres and Cuiabá identified as members of the species A. kuhlmannii, A. aff. kuhlmannii, and A. helodes (V 6325, V 10470, and Co 6862). Some regional overlap occurred between groups 1 and 2; for example, group 1 accession V 6352 and group 2 accession V 6351 were collected from neighboring sites.
Figure 5: Molecular characterization of Arachis kuhlmannii accessions and related species. Red, accessions collected in the region between Cáceres and Vila Bela da Santíssima Trindade; blue, accessions collected in the region between Cuiabá and Cáceres; green, accessions collected in Mato Grosso do Sul; violet, A. hypogaea; light blue, A. cardenasii; orange, A. magna and related accessions; black, cultivated accessions.
Group 3 included accessions collected in Mato Grosso do Sul, in the Miranda, Corumbá, and Aquidauana regions (Brazil). Within this group, genetic similarity closely correlates with geographic site of collection. The genetic similarity between groups 1, 2, and 3 was 0.52. As the farthest accessions within group 3 displayed an intragroup genetic similarity of 0.55, group 1, 2, and 3 accessions could belong to the same species.
Accessions EkZb 1 and V 13770 (plants b and c), initially identified as A. kuhlmannii, were transported from unknown locations to urban areas where they were grown in gardens. Accession V 13770 was collected at the Cáceres airport garden, MT, and accession EkZb 1 was collected from an ornamental lawn in the municipality of Lucas do Rio Verde, MT. A definitive identification of these accessions and their classification as A. kuhlmannii would require crosses with the typical accession KG 30034 or with the possible topotype V 13779.
Arachis cardenasii accession GKP 10017 remained isolated from other analyzed materials, displaying a genetic similarity of 0.45 with groups 1, 2, and 3 and accessions EkZb 1 and V 13770. We believe that this degree of similarity establishes the point of divergence between two species for the materials assessed here. Altogether, these findings support the classification proposed by Krapovickas and Gregory, who maintained A. cardenasii as a distinct species. Singh et al. (1991) also defended the separation of A. cardenasii from the other A-genome species when they stated that A. cardenasii and A. correntina have significantly different genomes in comparison to other species, indicating the existence of a new subgroup.
The accession of A. hypogaea ‘Tatu’ also remained isolated from the other groups mentioned above, which have diploid A-type genomes. The genetic similarity between ‘Tatu’ and other materials had a value of 0.36. Interestingly, our analysis placed the cultivated A. hypogaea ‘Tatu’ between the A- and B-genome groups. This finding gives support to the idea that the groups on either side of ‘Tatu’ correlate with the A and B genomes of A. hypogaea.
Group 4 includes accessions lacking the A chromosome pair, except for the specific V 13770-a, which remained grouped with accessions related to A. magna even though it has the A pair. We believe that an error occurred during sample collection in the screen house for molecular characterization. Accessions V 13765, V 13760, and V 13761 belong to the species A. magna. The accession V 6389 (A. gregoryi) has a genetic similarity of 0.39 with accessions of A. magna. Crosses made by C.E. Simpson corroborated the distance between A. gregoryi and A. magna (KGSSc 30097) and yielded hybrids with pollen viability of 12.8 and 13.5% (C.E.Simpson, personal communication).
Wild species of Arachis usually exist in isolated populations with few plants, and collectors obtain only small numbers of seeds at each site. As a result of this pattern, and because of most Arachis species self-pollinated, accessions should have a high degree of homozygosity (Galgaro et al., 1997).
Arachis kuhlmannii accessions were highly heterogeneous, with large genetic distances even between accessions in the same dendrogram group. Moretzsohn et al. (2013) observed this heterogeneity, stating that accessions of A-genome species tended to cluster, except A. kuhlmannii, whose accessions were distributed throughout the different A-genome species clusters.
In a dendrogram generated by molecular markers before the current study, A. simpsonii and A. helodes appeared isolated from the accessions identified as A. kuhlmannii, much as was observed for A. cardenasii . However, here we place A. simpsonii and A. helodes into two different groups that remain close to accessions belonging to A. kuhlmannii or related species. This distribution indicates a high degree of genetic similarity among these three species and casts doubt on the current nomenclature.
Regarding polymorphisms, only four were characterized as specific: Y19 (396-bp DNA segment exclusive to accessions with the B genome); Y19 (507-bp band specific for accession V6389); Y19 (555-bp DNA segment exclusive to diploid A-genome accessions); and W01 (677-bp band observed in accessions collected in Mato Grosso do Sul). The most commonly observed polymorphisms included bands that grouped accessions with no relation to any specific region or species.
Results obtained from molecular marker analysis show that groups 1, 2, and 3 correspond to geographical collection sites (Figure 5). This correlation suggests that these accessions share a single common ancestor and that geographical isolation drove the process of speciation in these populations. We believe, however, that the isolation occurred recently and that each group cannot be considered a different species. Thus, we would expect crosses between groups to have high pollen viability percentages. If future studies support the hypothesis of conspecificity, the older name A. helodes should be used for accessions in the three groups.
RAPD analysis provided a useful tool for the differentiation of Arachis germplasm. Galgaro et al. (1998) also obtained differentiation among materials included in sections Extranervosae Krapov. & W.C. Greg., Caulorrhizae Krapov. & W.C. Greg., Heteranthae Krapov. & W.C. Greg., and Triseminatae Krapov. & W.C. Greg. of the genus Arachis using RAPD markers.
Comparison between markers
Molecular markers distinguished accessions and identified groups that accompanied geographic collection sites. There was less distortion in the generation of clusters with correlation coefficients above 0.90. Accumulated variation reached 90% with 51 markers computed. However, 90 polymorphic bands were generated, thereby increasing the confidence of the data.
Plant taxonomists make use of many types of information, most notably related to the morphology and anatomy of plants, but also chemical composition, number and type of chromosomes and other characteristics. However, there is a tacit agreement that species must be visually recognizable. While similarity between individuals of a species has a genetic basis, species are recognized and defined by their external form (Hoyt, 1992). Here, the dendrogram based on morphological characteristics discriminated only two isolated groups: accessions with the B genome and A. hypogaea. Accessions with the A genome remained grouped, without clear morphological distinctions among them. This lack of discrimination potential could be expected because of the similarity among A-genome species in the Arachis section that require dichotomous keys for precise identification.
Conclusion
• Principal component analysis of morphological descriptors distinguished species and genomes. A-genome diploid species remained grouped, as did accessions with the B genome. Arachis hypogaea remained isolated.
• RAPD markers distinguished accessions and identified groups that matched geographic collection areas.
• Cytogenetic characterization confirmed the number of chromosomes and presence or absence of the A chromosome pair in accessions.
• Further taxonomic review of some species in the Arachis section is needed, as we could not discriminate as distinct species all of the accessions identified as A. kuhlmannii, A. helodes , and A. simpsonii by using morphological, molecular, and cytogenetic markers.
Conflicts of interest
The authors declare no conflict of interest.
Acknowledgments
A.P. Fávero and J.F.M. Valls thank National Council for Scientific and Technological Development (CNPq/Brazil) for grants (#312215/2013-4, #483860/2012-3) and financial support.
About the Authors
Corresponding Author
A.P. Fávero
Embrapa Pecuária Sudeste, São Carlos, SP, Brasil
- Email:
- alessandra.favero@embrapa.br
References
- Bechara MD, Moretzsohn MC, Palmieri DA, Monteiro JP, et al. (2010). Phylogenetic relationships in genus Arachis based on ITS and 5.8S rDNA sequences. BMC Plant Biol. 10: 255. https://doi.org/10.1186/1471-2229-10-255
- Fávero AP, Moraes SA, Garcia AAF, Valls JFM, et al. (2009). Characterization of rust, early and late leaf spot resistance in wild and cultivated peanut germplasm. Sci. Agr. 66: 110-117. https://doi.org/10.1590/S0103-90162009000100015
- Fernández A and Krapovickas A (1994). Cromosomas y evolución en Arachis (Leguminosae). Bonplandia 8: 187-220.
- Ferreira ME and Grattapaglia D (1996). Introdução ao Uso de Marcadores Moleculares em Análise Genética. Embrapa CENARGEN, Brasília.
- Galgaro L, Valls JFM and Lopes CR (1997). Study of the genetic variability and similarity among and within Arachis villosulicarpa, A. pietrarelli and A. hypogaea through isoenzyme analysis. Genet. Resour. Crop Evol. 44: 9-15.https://doi.org/10.1023/A:1008609115357
- Galgaro L, Lopes CR, Gimenes MA, Valls JFM, et al. (1998). Genetic variation between several species of sections Extranervosae, Caulorrhizae, Heteranthae, and Triseminatae (genus Arachis) estimated by DNA polymorphism. Genome 41: 445-454.https://doi.org/10.1139/g98-004
- Gimenes MA, Hoshino AA, Barbosa AV, Palmieri DA, et al. (2007). Characterization and transferability of microsatellite markers of the cultivated peanut (Arachis hypogaea). BMC Plant Biol. 7: 9. https://doi.org/10.1186/1471-2229-7-9
- Gregory WC, Krapovickas A and Gregory MP (1980). Structure, variation, evolution and classification in Arachis. In: Advances in Legume Science (Summerfield RJ and Bunting AH, eds.), Kew, London, 2: 469-481.
- Hoyt E (1992). Conservação dos Parentes Silvestres das Plantas Cultivadas. IBPGR, IUCN e WWF.
- Hussaini SH, Goodman MM and Timothy DH (1977). Multivariate analysis and geographical distribution of the world collection of finger millet. Crop Sci. 17: 257-263. https://doi.org/10.2135/cropsci1977.0011183X001700020007x
- IBPGR (1990). Preliminary Descriptors for Arachis (IBPGR/ICRISAT) - International Crop Network Series. 2. Report of a Workshop on the Genetic Resources of Wild Arachis Species. International Board for Plant Genetic Resources, Rome.
- Kornerup A and Wanscher JH (1979). Methuen Handbook of Colors. 3 ed. Methuen, London.
- Krapovickas A and Gregory WC (1994). Taxonomía del género Arachis (Leguminosae). Bonplandia 8: 1-186.
- Lodhi AM, Ye GN, Weeden NF and Reisch BI (1994). A simple and efficient method for DNA extraction from grapevine cultivars and Vitis species. Plant Mol. Biol. Rep. 12: 6-13. https://doi.org/10.1007/BF02668658
- Michelotto MD, Godoy IJ, Fávero AP, Carrega WC, et al. (2013). Occurrence of Enneothrips flavens Moulton and Stegasta bosquella (Chambers) and its effects on agronomic traits of wild Arachis accessions. Biosci. J. 29: 115-124.
- Monçato L (1995). Caracterização morfológica de germoplasma de espécies de Arachis, secção Caulorrhizae, pela análise multivariada. Botucatu. MSc dissertation, Instituto de Biociências, Universidade Estadual Paulista Júlio Mesquita Filho.
- Moretzsohn MC, Gouvea EG, Inglis PW, Leal-Bertioli SCM, et al. (2013). A study of the relationships of cultivated peanut (Arachis hypogaea) and its most closely related wild species using intron sequences and microsatellite markers. Ann. Bot. 111: 113-126.https://doi.org/10.1093/aob/mcs237
- Pott A, Oliveira AKM, Damasceno-Junior GA and Silva JSV (2011). Plant diversity of the Pantanal wetland. Braz. J. Biol. 71 (Suppl 1): 265-273. https://doi.org/10.1590/S1519-69842011000200005
- Robledo G, Lavia GI and Seijo G (2009). Species relations among wild Arachis species with the A genome as revealed by FISH mapping of rDNA loci and heterochromatin detection. Theor. Appl. Genet. 118: 1295-1307. https://doi. org/10.1007/s00122-009-0981-x
- Singh AK, Sivaramakrishnan S, Mengesha MH and Ramaiah CD (1991). Phylogenetic relations in section Arachis based on seed protein profile. Theor. Appl. Genet. 82: 593-597. https://doi.org/10.1007/BF00226795
- Tallury SP, Hilu KW, Milla SR, Friend SA, et al. (2005). Genomic affinities in Arachis section Arachis (Fabaceae): molecular and cytogenetic evidence. Theor. Appl. Genet. 111: 1229-1237. https://doi.org/10.1007/s00122-005-0017-0
- Valls JFM, Simpson CE and Ramanatha R (1995). Collecting wild species of Arachis. In: Collecting Plant Genetic Diversity (Guarino L, Rao VR and Reid R, eds.). Technical Guidelines. CAB International Chapter, Wallingford, 35: 677-684.
- Williams JGK, Kubelik AR, Livak KJ, Rafalski JA, et al. (1990). DNA polymorphisms amplified by arbitrary primers are useful as genetic markers. Nucleic Acids Res. 18: 6531-6535. https://doi.org/10.1093/nar/18.22.6531
Keywords:
Download:
Full PDF- Share This