Molecular insights of mitochondrial 16S rDNA genes of the native honey bees subspecies Apis mellifera carnica and Apis mellifera jementica (Hymenoptera: Apidae) in Saudi Arabia
Received: November 30, 2018
Accepted: December 21, 2018
Published: January 05, 2019
Genet.Mol.Res. 18(3):
Introduction
Bees are the world’s most important pollinators in natural and agricultural ecosystems. They are very diverse with over 20,000 species (Engel, 2005, 2011a). Honey bees belong to the family Apidae with other social bees. The subfamily Apidae is comprised of a single genus, Apis . The genus Apis comprises Apis florae (the little bee), Apis dorsata (the giant bee), Apis cerana (the eastern bee) and Apis mellifera (the western bee). The honey bee, Apis mellifera Linnaeus, 1758, is naturally found throughout Europe, Africa and Western Asia (Miguel et al., 2011). Traditionally, the intraspecific taxonomy of the honey bee Apis mellifera L. has been based on morphology. Twenty-nine subspecies of A. mellifera are currently recognized on the basis of morphometric characteristics (Sheppard and Meixner, 2003; Miguel et al., 2011). These subspecies are also described as ‘geographic races’ because their distributions correspond to distinct geographic areas (Ruttner, 1992). Five evolutionary lineages have been characterized based on morphometric, molecular, ecological, ethological, and physiological traits (De la Rùa et al., 2005). The four primary lineages are found in the Mediterranean Basin, including the African lineage (A), west and north European lineage (M), south-east European lineage (C), and Near and Middle Eastern lineage (O) (Franck et al., 2000a, 2001; Miguel et al., 2007; Cánovas et al., 2008). The fifth proposed lineage is north-east African (Y) (Franck et al., 2001). Although morphological characteristics are still considered very important in the classification of honey bees, this approach is not well suited to characterize honey bee subspecies and analyze phylogenetic relationships because they can be sensitive to environmental selection pressures (Franck et al., 2000b).
A number of molecular markers such as nuclear DNA (Hall, 1990; Tarès et al., 1993), mitochondrial DNA (mtDNA) (Moritz et al., 1986; Smith et al., 1989; Smith, 1991; Hunt and Page, 1992; Garnery et al., 1993; Oldroyd et al., 1995; Arias and Sheppard, 1996; Pedersen, 1996; De la Rúa et al., 2000), and microsatellites (Estoup et al., 1993; Garnery et al., 1998), are also used to study genetic variability in honey bees (Boore, 1999). The sequencing and characterization of the mtDNA genome have been very useful for analyzing the phylogeny and population genetic structure of the Apis species and of A. mellifera subspecies because it contains regions with variable evolutionary rates. In general, mtDNA contains genes for two ribosomal subunits (12S and 16S), 22 tRNA, and 13 proteins (three subunits of cytochrome c oxidase, cytochrome B, subunits 6 and 8 of ATP F0 synthase, and seven subunits of NADH dehydrogenase) (Shao et al., 2003; Silvestre and Arias, 2006). Genetic markers such as the mtDNA COXI–COXII intergenic region are unique to the genus Apis (Cornuet and Garnery 1991). Variations in the sequences of this region or the length of fragments produced using endonucleases are used extensively to differentiate the five honeybee lineages and to discriminate among A. mellifera subspecies (Garnery et al., 1992; Franck et al., 2000a; Sheppard and Smith, 2000). To date, no study has characterized Saudi honey bees. Therefore, the goal of this study was to determine the genetic diversity and phylogenetic relationships of Apis mellifera subspecies of Saudi Arabia via mitochondrial 16S rDNA genes.
Material and Methods
Honey bee sample collection
A total of 100 samples were collected from adult worker bees of each breeder (200 samples) from honey bees in the Hail region of Saudi Arabia. Bee samples were collected during the month of September 2017 after approval from the beekeepers. Bees were collected inside the wooden cages containing food with passage of air and sunlight. The samples were transferred alive to an Entomology lab, frozen for 5 min to immobilize them, and then stored in 95% ethanol until processed. The two strains were identified according to the external characteristics with measurements of different body parts. Some samples were sent to a specialist for definition of the bee breeds with Dr. Yahya Al-Attal, Faculty of Agriculture, King Saud University, Saudi Arabia.
Molecular analysis
DNA extraction, PCR amplification, and sequencing: Mitochondrial genomic DNA was extracted from the thoracic legs of the ethanol-preserved samples. The tissue was homogenized using a glass pestle and a mortar. The genomic DNA in the homogenate was extracted using DNeasy Blood and Tissue Kit (Qiagen, Germany) following the manufacturer's instructions. The purity and concentration of extracted DNA was determined with a NanoDrop 2000 UV-VIS Spectrophotometer (Thermo Fisher Scientific Inc, USA) at 260/280 nm. The extracted DNA was stored at -20°C until use. Mitochondrial 16S rDNA gene fragments were amplified using GeneJETTM PCR Purification kit [Thermo (Fermentas)] following the manufacturer's protocol in a total volume of 50 μl including 5 μl 10 × buffer, 5 μl of each dNTP (10 mM), 10 μl of each primer (1 pmol/ μl), 0.3 μl of Taq polymerase (5 U/ml), 2.5 μl MgCl2 (50 mM), and 2 μl of total genomic DNA. PCR amplification and subsequent DNA sequencing was carried out using the following mitochondrial 16S rDNA primers: 16SF (5’-CAC CTG TTT ATC AAA AAC AT-3’) and 16SR (5’-CGT CGA TTT GAA CTC AAA TC-3’), designed by Crozir and Crozier (1993).
The PCR cyclic conditions were processed under the following conditions: 95°C for 15 min (initial denaturation), 35 cycles of 15 s at 95°C (denaturation), 30 s at 62°C (annealing), and 1 min at 72°C (extension), and finally post-PCR extension for 5 min at 72°C. Each amplicons was examined by (1%) agarose gel electrophoresis in 1× Tris–acetate–EDTA (TAE) buffered gel stained with 1% Ethidium bromide. This was then visualized with a UV transilluminator, and bands with the predicted size were purified using QIAquick PCR Purification Kit (Qiagen) following the manufacturer’s instructions. Amplicons were sequenced (in both directions) using an ABI Prism Dye Terminator Cycle Sequencing Core Kit (Applied Biosystems; Thermo Fisher Scientific, Waltham, MA, USA) with an Applid Analyzer Biosystems (Thermo Fisher Scientific, USA) using the same primers for annealing.
Sequence alignment and phylogenetic analysis
BLAST search was done to identify related sequences on NCBI database. Sequences were aligned directly using CLUSTAL-X multiple sequence alignment (Thompson et al., 1997) and others available from GenBankTM selected to represent all available tapeworm lineages with an emphasis on taxa presumed to be related to the analyzed groups. The alignment was manually corrected using the alignment editor of software BIOEDIT 4.8.9 (Hall, 1999). Phylogenetic calculations were performed with the UPGMA method. The data were analyzed with the Maximum Composite Likelihood (MCL) approach based on the Kimura 2-parameter model. The percentage of replicate trees in which the associated taxa clustered together in the bootstrap test (1000 replicates) are shown next to the branches. Phylogenetic and evolutionary analyses were conducted using MEGA version 6 (Tamura et al., 2013). The tree was drawn to scale with branch lengths in the same units as those of the evolutionary distances used to infer the phylogenetic tree. The evolutionary distances were computed using the p-distance method (Nei and Kumar, 2000).
Results
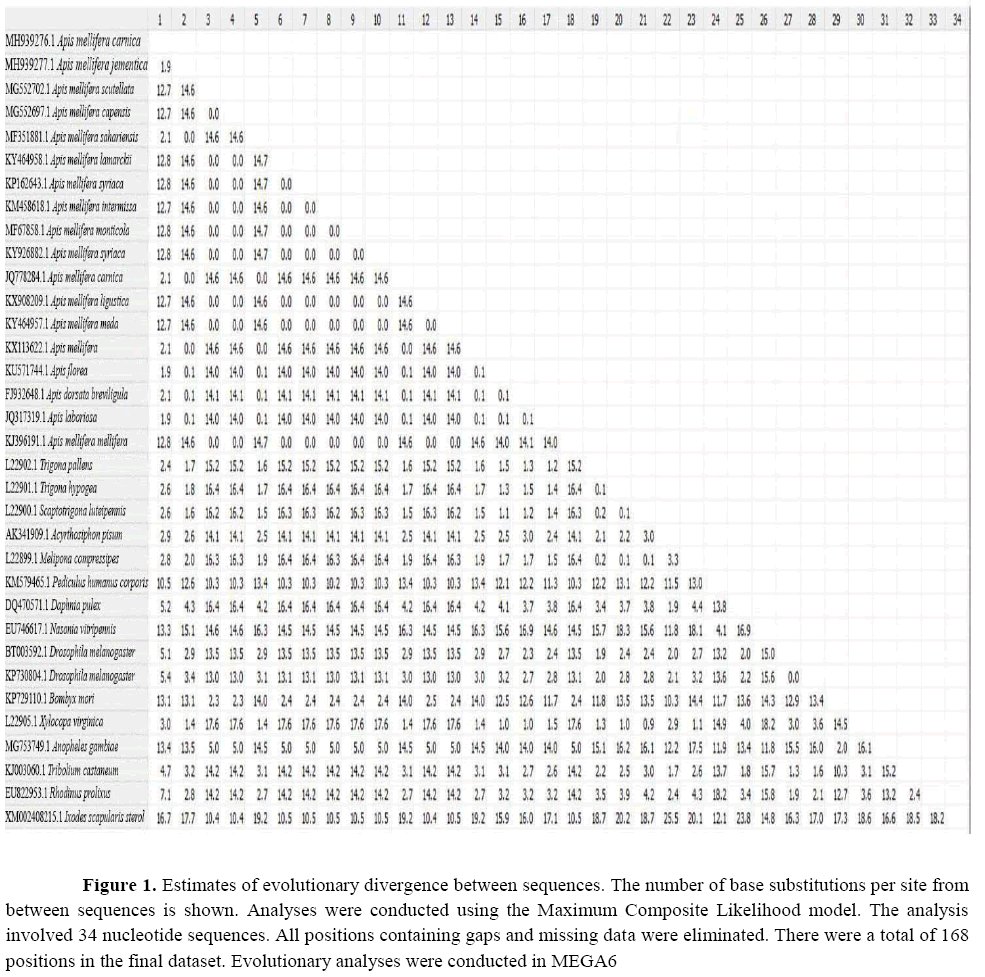
Molecular analyses based on the partial 16S rDNA gene sequence were performed to investigate the taxonomy and classification of the two recovered Apis mellifera subspecies. The amplified and sequenced gene regions and GC content in the samples from Apis mellifera carnica and Apis mellifera jementica were 1157 bp/29% and 682 bp/25.7%, respectively. The sequences were deposited in GenBank under the accession numbers, MH939276.1 and MH939277.1, respectively. The sequences were compared with each other and with other sequences available on GenBank from varied geographical regions (Table 1). The calculation of the percentage of identity between these novel sequences and a range of sequences for other arthropods of different host species demonstrated a high degree of similarity (up to 72%) with intra-specific differences that varied from 0.1% to 23.8% (Figure 1).
| Parasite species | Class/Order/Family | Source | Accession no. | Sequence length (bp) | Percent identity (%) |
|---|---|---|---|---|---|
| Apis mellifera scutellata | Insecta/Hymenoptera/Apidae | GenBank | MG552702.1 | 16364 | 91 |
| Apis mellifera capensis | Insecta/Hymenoptera/Apidae | GenBank | MG552697.1 | 16456 | 91 |
| Apis mellifera sahariensis | Insecta/Hymenoptera/Apidae | GenBank | MF351881.1 | 16569 | 91 |
| Apis mellifera lamarckii | Insecta/Hymenoptera/Apidae | GenBank | KY464958.1 | 16589 | 91 |
| Apis mellifera syriaca | Insecta/Hymenoptera/Apidae | GenBank | KP162643.1 | 15428 | 91 |
| Apis mellifera intermissa | Insecta/Hymenoptera/Apidae | GenBank | KM458618.1 | 16343 | 91 |
| Apis mellifera monticola | Insecta/Hymenoptera/Apidae | GenBank | MF67858.1 | 16343 | 91 |
| Apis mellifera syriaca | Insecta/Hymenoptera/Apidae | GenBank | KY926882.1 | 16343 | 91 |
| Apis mellifera carnica | Insecta/Hymenoptera/Apidae | GenBank | JQ778284.1 | 1009 | 91 |
| Apis mellifera ligustica | Insecta/Hymenoptera/Apidae | GenBank | KX908209.1 | 16343 | 91 |
| Apis meellifera meda | Insecta/Hymenoptera/Apidae | GenBank | KY464957.1 | 16248 | 91 |
| Apis mellifera | Insecta/Hymenoptera/Apidae | GenBank | KX113622.1 | 515 | 91 |
| Apis florea | Insecta/Hymenoptera/Apidae | GenBank | KU571744.1 | 520 | 90 |
| Apis dorsata breviligula | Insecta/Hymenoptera/Apidae | GenBank | FJ932648.1 | 468 | 90 |
| Apis laboriosa | Insecta/Hymenoptera/Apidae | GenBank | JQ317319.1 | 469 | 90 |
| Apis mellifera mellifera | Insecta/Hymenoptera/Apidae | GenBank | KJ396191.1 | 16051 | 91 |
| Trigona pallens | Insecta/Hymenoptera/Apidae | GenBank | L22902.1 | 458 | 76 |
| Trigona hypogea | Insecta/Hymenoptera/Apidae | GenBank | L22901.1 | 459 | 79 |
| Scaptotrigona luteipennis | Insecta/Hymenoptera/Apidae | GenBank | L22900.1 | 461 | 78 |
| Acyrthosiphon pisum | Insecta/Homoptera/Aphididae | GenBank | AK341909.1 | 661 | 74 |
| Melipona compressipes | Insecta/Hymenoptera/Apidae | GenBank | L22899.1 | 459 | 76 |
| Pediculus humanus corporis | Insecta/Phthirapter/Pediculidae | GenBank | KM579465.1 | 668 | 82 |
| Daphnia pulex | Crustacea/Branchiopoda/Daphniidae | GenBank | DQ470571.1 | 442 | 72 |
| Nasonia vitripennis | Insecta/Hymenoptera/Pteromalidae | GenBank | EU746617.1 | 2747 | 78 |
| Drosophila melanogaster | Insecta/Diptera/Drosophilidae | GenBank | BT003592.1 | 3412 | 72 |
| Drosophila melanogaster | Insecta/Diptera/Drosophilidae | GenBank | KP730804.1 | 342 | 73 |
| Bombyx mori | Insecta/Lepidoptera/Bombycidae | GenBank | KP729110.1 | 15658 | 77 |
| Xylocopa virginica | Insecta/Hymenoptera/Apidae | GenBank | L22905.1 | 463 | 82 |
| Anopheles gambiae | Insecta/Diptera/Culicidae | GenBank | MG753749.1 | 14844 | 74 |
| Tribolium castaneum | Insecta/Coleoptera/Tenebrionidae | GenBank | KJ003060.1 | 526 | 72 |
| Rhodinus prolixus | Insecta/Hemiptera/Reduviidae | GenBank | EU822953.1 | 316 | 72 |
| Ixodes scapularis sterol | Chelicerata/Arachnida/Ixodidae | GenBank | XM002408215.1 | 3339 | 72 |
Table 1. Arthropoda species used in the phylogenetic analysis of the present sub-species Apis mellifera
Figure 1: Estimates of evolutionary divergence between sequences. The number of base substitutions per site from between sequences is shown. Analyses were conducted using the Maximum Composite Likelihood model. The analysis involved 34 nucleotide sequences. All positions containing gaps and missing data were eliminated. There were a total of 168 positions in the final dataset. Evolutionary analyses were conducted in MEGA6
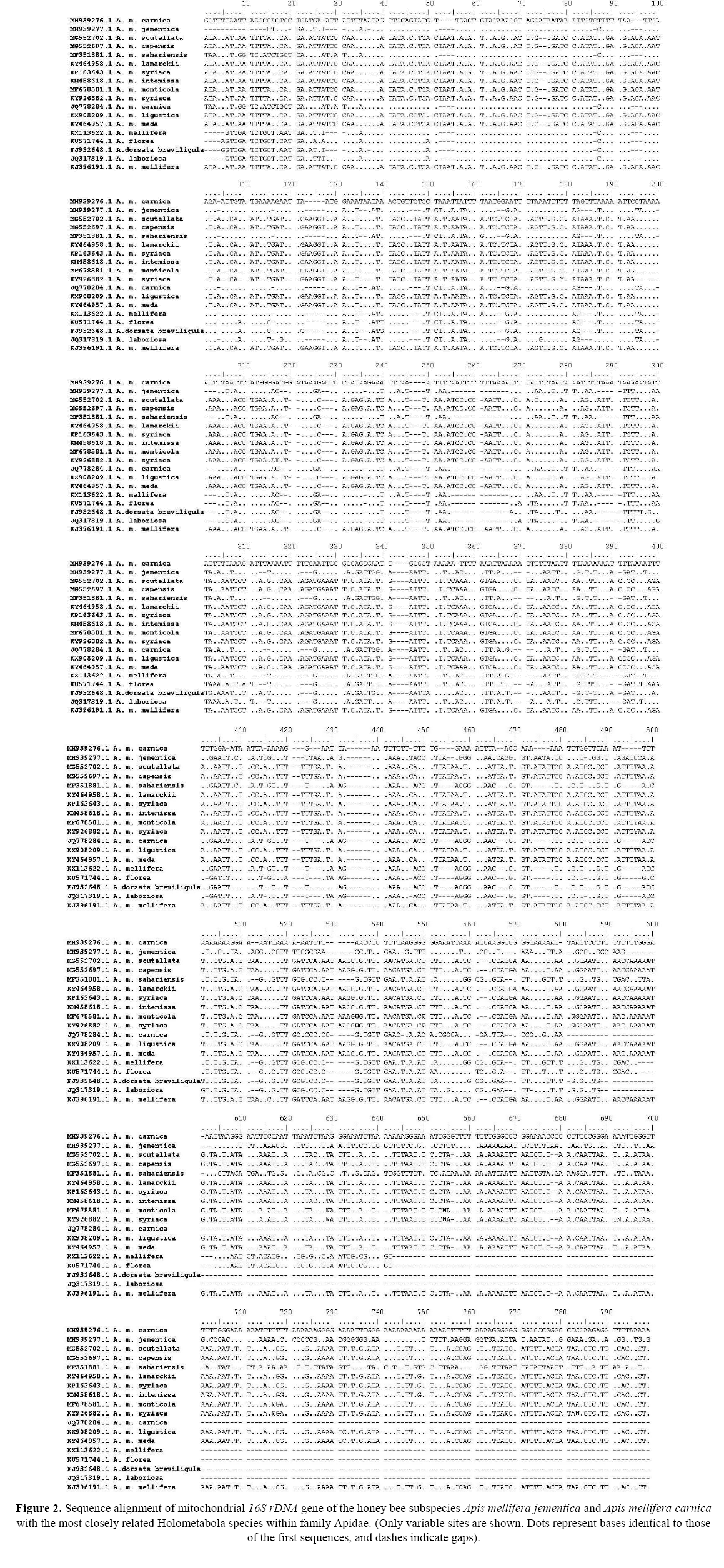
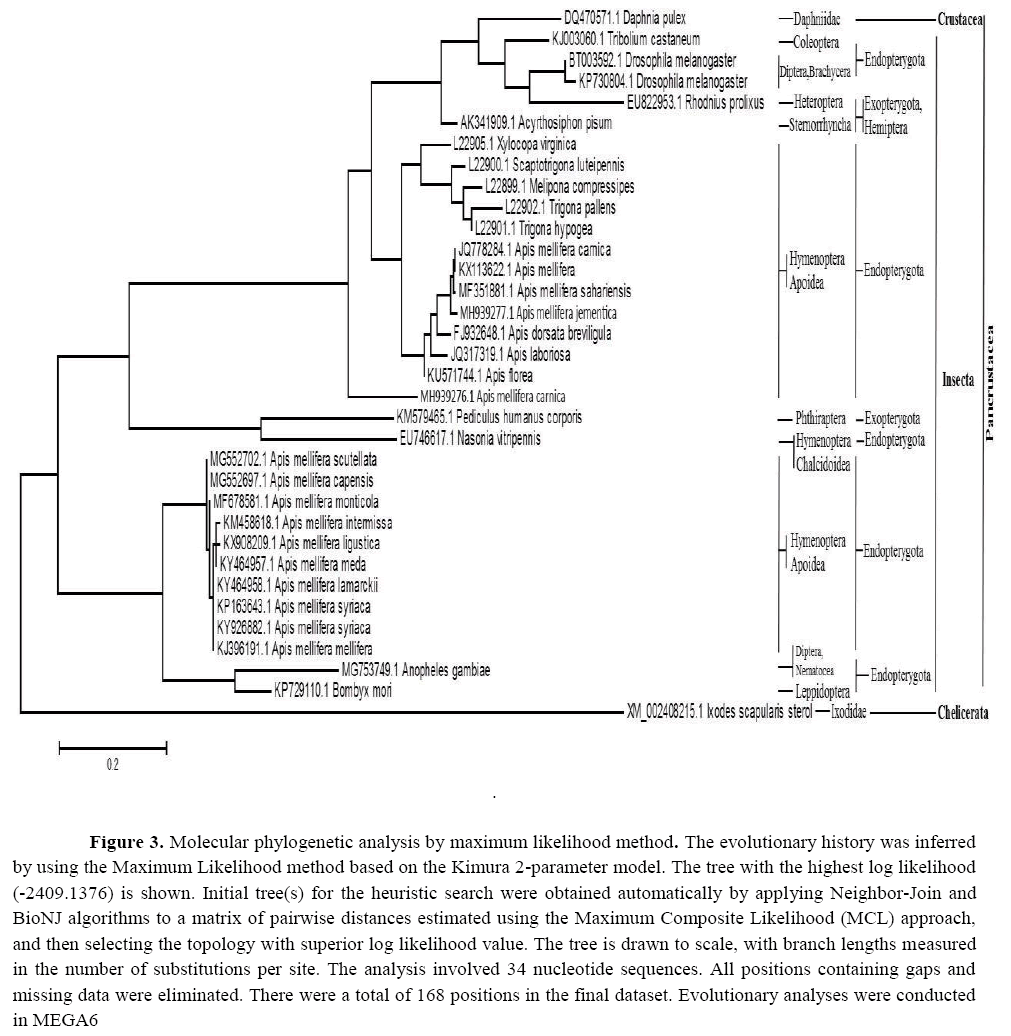
The comparison of nucleotide sequences and divergence showed that 16S rDNA in the present species has the highest BLAST scores with lower divergence values than other apids previously sequenced; however, our sample was not identical to any prior specimen (Figure 2). A tree topology was automatically computed to estimate ML values (Figure 3). Two clades were clustered during the construction of the phylogenetic tree using the maximum likelihood and maximum parsimony of the present apids species. The major clade clustering all Pancrustacean species contained two classes: the first one was Crustacea represented here by Daphnia pulex (gb| DQ470571.1) within family Daphniidae; the second was Insecta with families Apidae ( A. mellifera scutellata gb| MG552702.1, A. mellifera capensis gb| MG552697.1, A. mellifera sahariensis gb| MF351881.1, A. mellifera Lamarckii gb| KY464958.1, A. mellifera syriaca gb| KP162643.1, A. mellifera intermissa gb| KM458618.1, A. mellifera monticola gb| MF67858.1, A. mellifera syriaca gb| KY926882.1, A. mellifera carnica gb| JQ778284.1, A. mellifera ligustica gb| KX908209.1, A. meellifera meda gb| KY464957.1, A. mellifera gb| KX113622.1, A. florea gb| KU571744.1, A. dorsata breviligula gb| FJ932648.1, A. laboriosa gb| JQ317319.1, A. mellifera mellifera gb| KJ396191.1, Trigona pallens gb| L22902.1, T. hypogea gb| L22901.1, Scaptotrigona luteipennis gb| L22900.1, Melipona compressipes gb| L22899.1, Xylocopa virginica gb| L22905.1), Aphididae (Acyrthosiphon pisum gb| AK341909.1), Pediculidae (Pediculus humanus corporis gb| KM579465.1), Pteromalidae (Nasonia vitripennis gb| EU746617.1), Drosophilidae (Drosophila melanogaster gb| BT003592.1, gb| KP730804.1), Bombycidae (Bombyx mori gb| KP729110.1), Cuclicidae (Anopheles gambiae gb| MG753749.1), Tenebrionidae (Tribolium castaneum gb| KJ003060.1), and Reduviidae (Rhodinus prolixus gb| EU822953.1). The minor clade with the lowest blast scores and a high divergence value contained class Chelicerata represented by Ixodes scapularis sterol (gb| XM002408215.1). These results indicate that the recovered apid species are deeply embedded within the genus Apis .
Figure 2: Sequence alignment of mitochondrial 16S rDNA gene of the honey bee subspecies Apis mellifera jementica and Apis mellifera carnica with the most closely related Holometabola species within family Apidae. (Only variable sites are shown. Dots represent bases identical to those of the first sequences, and dashes indicate gaps).
Figure 3: Molecular phylogenetic analysis by maximum likelihood method. The evolutionary history was inferred by using the Maximum Likelihood method based on the Kimura 2-parameter model. The tree with the highest log likelihood (-2409.1376) is shown. Initial tree(s) for the heuristic search were obtained automatically by applying Neighbor-Join and BioNJ algorithms to a matrix of pairwise distances estimated using the Maximum Composite Likelihood (MCL) approach, and then selecting the topology with superior log likelihood value. The tree is drawn to scale, with branch lengths measured in the number of substitutions per site. The analysis involved 34 nucleotide sequences. All positions containing gaps and missing data were eliminated. There were a total of 168 positions in the final dataset. Evolutionary analyses were conducted in MEGA6
Discussion
Apidae is the largest family of bees, with over 5,700 described species (Ascher and Pickering, 2012). There are four different species of honey bees in the world: little honey bee (Apis florea) that is native to southeast Asia, the Eastern honey bee (Apis cerana) of eastern Asia including Korea and Japan; the Giant honey bee (Apis dorsata) native to southeast Asia; and the Western honey bee (Apis mellifera) native to Europe, Africa, and western Asia (Daly et al., 1982; Oldroyd and Wongsiri, 2006; Rattanawannee et al., 2010). Apis mellifera is commonly known as the European, western, or common honeybee in different parts of the world and is extensively domesticated for honey, pollen, beeswax, propolis, royal jelly, and bee venom (Ratcliffe et al., 2011). It is a major agent of global ecology via pollination (Williams, 2002; Kajobe, 2006; Klein et al., 2007; Allsopp et al., 2008). It is a common model in scientific social behavioral studies (Breed et al., 2004; Solignac et al., 2004; Beye et al., 2006; The Honey Bee Genome Sequencing Consortium, 2006). The current study aimed to determine the genetic diversity and phylogenetic relationships of the A. mellifera subspecies of Saudi Arabia. The Apis species are often identified via molecular markers due to the lack of taxonomists skilled in apiculture in Saudi Arabia.
DNA sequence-based identification techniques have revealed the morphological and ecological traits of many species during larval stages (Foltan et al., 2005; Smith et al., 2006; Hayashi and Sota, 2010). Mitochondrial DNA (mtDNA) markers have been widely used to address population and evolutionary questions from the A. mellifera honey bee. The sequencing and characterization of the mitochondrial DNA genome has been very useful for analyzing the phylogeny and population genetic structure of the Apis species and of the A. mellifera subspecies. It contains regions with variable evolutionary rates. In principle, the general pattern of subspecies distribution has been supported by various genetic studies using molecular tools (Garnery et al., 1993; Franck et al., 2000a; 2001; Whitfield et al., 2006; Cánovas et al., 2008). Mitochondrial introgressions can be assessed using the “DraI” test-a molecular test that highlights the sequence variability between subunits I and II of the cytochrome oxidase gene (Garnery et al., 1993). This test has been widely used to analyze the biogeography of A. mellifera L. subspecies and races (Rortais et al., 2011) as well as for other Apis species (Smith and Hagen, 1997).
We used this method to distinguish different haplotypes and group them into one of the four primary lineages (Ruttner, 1988; De la Rúa et al., 2009). The mitochondrial 16S rDNA was studied here for the first time to detect the molecular polymorphism between two different subspecies of Apis mellifera worker honey bee. Previously, the mitochondrial CoxI–CoxII intergenic region was utilized to distinguish honey bee lineages. Schiff and Sheppard (1995) used mitochondrial DNA and allozyme variations to characterize 142 breeder queen colonies from 22 apiaries in the southeastern United States. In addition, Franck et al. (2000a) described the different haplotypes of the CoxI-CoxII intergenic region found in major honey bee lineages. Kozmus et al. (2007) analyzed the mtDNA of honeybee ectotypes as the initial step of molecular characterization of the indigenous honeybee populations from Serbia. Papachristoforou et al. (2013) combined analysis of mtDNA and microsatellite data from A. m. cypria to describe the genetic structure of the Cyprian honey bee population. Techer et al. (2015) analyzed the genetic diversity of the honeybee population in Rodrigues using the mitochondrial CoxI-CoxII intergenic region. Bo�?¾i�? et al. (2016) analyzed mtDNA for the characterization of carniolan honey bee A. M. carnica.
This study indicated that closely related Apis species have 90% similarity in the standardized DNA sequence and distantly related species have less than 90% similarity in the same genes sequence. This agrees with Gurney et al. (2000). The NJ tree was constructed here based on the multiple aligned sequence data for different Apis species and other arthropods. The tree separates the genomes into two main clades. All A. mellifera species were included in one clade; Chelicerata represented by Ixodes scapularis sterol were in another clade. Our results verify the conclusions of Prentice (1998), Melo (1999), Michener (2007), Cardinal et al. (2010), and Haddad et al. (2017) who found that all Apis species are clustered together in one clade in addition to the genetic origin of Apis species within family Apidae as a paraphyletic group.
Franck et al. (2001) and Cánovas et al. (2008) demonstrated that there are five honey bee lineages found in the Mediterranean Basin. These results indicated that the recovered apid species are deeply embedded within the genus Apis within the African lineage. A recent similar study by Haddad et al. (2017) analyzed the mt genome haplotyes and indicated that the north African Sahara Honey bee, A. m. sahariensis in the African lineage was genetically distinct from the northern African lineage exemplified by A. m. intermissa. The branch length of A. m. carnica (gb| MH939276.1) was lower than A. m. jementica (gb| MH939277.1) indicating less divergence from its ancestor. This result was confirmed by data from Rukhsana et al. (2014) who indicated that the branch length of the NJ tree is a degree to indicate the divergence from the common ancestor.
Conclusion
A recent field study provides tools for the rapid identification and phylogenetic analysis of honey bees inhabiting Saudi Arabia via mitochondrial 16S rDNA gene from the recovered Apis mellifera subspecies. This yielded a unique genetic sequence that confirms their taxonomic position within the family Apidae.
Competing Interests
The authors have declared that they have no conflict of interest regarding the content of this article.
Acknowledgment
This research project supported by a grant from the Research Center of the Female Scientific and Medical Colleges, Deanship of Scientific Research, King Saud University, Saudi Arabia. We thank the Deanship of Scientific Research and RSSU at King Saud University for their technical support.
COMPLIANCE WITH ETHICAL STANDARDS
All procedures contributing to this work comply with the ethical standards of the relevant national guides on the are and use of laboratory animals and have been approved and authorized by the Institutional Animal Care and Use Committee (IACUC) at King Saud University, Riyadh, Saudi Arabia.
About the Authors
Corresponding Author
Rewaida Abdel-Gaber
Department of Zoology, College of Science, King Saud University, Riyadh, Saudi Arabia
- Email:
- rewaida@sci.cu.edu.eg
References
- Al-Ghamdi AA (1990) Survey of honeybee diseases, pests and predators in Saudi Arabia. MPhil Thesis, University of Wales, Cardiff, United Kingdom, pp 17-127.
- Alattal Y and Al-Ghamdi A (2015) Impact of temperature extremes on survival of indigenous and exotic honey bee subspecies, Apis mellifera, under desert and semiarid climates. B Insectol 68: 219–222.
- AlGhamdi A, Al-Sharhi M, AlAttal Y, Adgaba N (2012) Morphometric diversity of indigenous honey bees, Apis mellifera (Linnaeus, 1758), in Saudi Arabia. Zoology in the Middle East 58: 97-103. https://doi.org/10.1080/09397140.2012.10648968
- Al-Ghamdi AA (2010). Comprehensive Study for Current Beekeeping Industry of Imported and Native Honey bee in Kingdom of Saudi Arabia. King Saud University, College of Agriculture, Bee Research Unit, Riyadh.
- Allsopp MH, de Lange WJ, Veldtman R (2008) Valuing Insect Pollination Services with Cost of Replacement. PLoS ONE 3: e3128. https://doi.org/10.1371/journal.pone.0003128
- Ansari MJ, Al-Ghamdi A, Nuru A, Ahmed AM, et al. (2016) Survey and molecular detection of Melissococcus plutonius, the causative agent of European Foulbrood in honey bees in Saudi Arabia. Saudi J Biol Sci 24 : 1327-1335. https://doi.org/10.1016/j.sjbs.2016.10.012
- Ansari MJ, Al-Ghamdi A, Usmani S, Khan KA, et al. (2017) In vitro evaluation of the effects of some plant essential oils on Ascosphaera apis, the causative agent of Chalkbrood disease. Saudi J Biol Sci 24 : 1004–1011. https://doi.org/10.1016/j.sjbs.2016.04.016
- Arias MC and Sheppard WS (1996) Molecular phylogenetics of honey bee subspecies (Apis mellifera L.) inferred from mitochondrial DNA sequence. Molecular Phylogenetics and Evolution 5: 557-566. https://doi.org/10.1006/mpev.1996.0050
- Ascher JS and Pickering J (2012) Discover life: bee species guide and world checklist (Hymenoptera: Apoidea: Anthophila).
- Beye M, Gattermeier I, Hasselmann M, Gemp T, et al. (2006) Exceptionally high levels of recombination across the honeybee genome. Genome Research 16: 1339–1344. https://doi.org/10.1101/gr.5680406
- Boore J (1999) Animal mitochondrial genomes. Nucleic Acids Res 27: 1767-1780. https://doi.org/10.1093/nar/27.8.1767
- Bo�?¾ic J, Kordiš D, Kri�?¾aj I, Leonardi A, et al. (2016) Novel aspects in characterisation of carniolan honeybee (Apis mellifera carnica, Pollmann 1879). Animal Science Days 5: 18-27.
- Breed MD, Guzman-Novoa E, Hunt GJ (2004) Defensive Behavior of Honey Bees: Organization, Genetics, and Comparisons with Other Bees. Annual Reviews of Entomology 49: 271–298. https://doi.org/10.1146/annurev.ento.49.061802.123155
- Cánovas F, De la Rúa P, Serrano J, Galián J (2008) Geographical patterns of mitochondrial DNA variation in Apis mellifera iberiensis (Hymenoptera: Apidae). Journal of Zoological Systematics and Evolutionary Research 46: 24-30. https://doi.org/10.1111/j.1439-0469.2007.00435.x
- Cardinal S, Straka J, Danforth BN (2010) Comprehensive phylogeny of apid bees reveals the evolutionary origins and antiquity of cleptoparasitism. Proc. Natl. Acad. Sci. USA 107: 16207–16211. https://doi.org/10.1073/pnas.1006299107
- Cornuet J and Garnery L (1991) Mitochondrial DNA variability in honey bees and its phylogeographic implications. Apidologie 22: 627-642. https://doi.org/10.1051/apido:19910606
- Crailsheim K, Brodschneider R, Neumann P (2009) The COLOSS puzzle: filling in the gaps. In: Proceedings of the 4th COLOSS Conference, 3-4 March 2009. Zagreb, Croatia, p. 46-47.
- Crozier RH and Crozier YC (1993) The mitochondri¬al genome of the honeybee Apis mellifera: complete sequence and genome organization. Genetics 133: 97-117.
- Daly HV, Hoelmer K, Norman P, Allen T (1982) Computer assisted measurement and identification of honey bees. Annals of Entomological Society of America 75: 591-594. https://doi.org/10.1093/aesa/75.6.591
- De la Rúa P, Fuchs S, Serrano J (2005) Biogeography of European honeybees. In: Lodesani M., Costa C. (Eds.) Beekeeping and Conserving Biodiversity of Honeybees. Sustainable Bee Breeding. Theoretical and practical guide. Northern Bee Books. Hebden Bridge 15-52.
- De la Rúa P, Jaffé R, Dall’Olio R, Muños I, et al. (2009) Biodiversity, conservation and current threats to European honeybees. Apidologie 40: 263-284. https://doi.org/10.1051/apido/2009027
- De la Rúa P, Simon UE, Tide AC, Moritz RFA, et al. (2000) MtDNA variation in Apis cerana populations from the Philippines. Heredity 84: 124-130. https://doi.org/10.1046/j.1365-2540.2000.00646.x
- Dowton M, Campbell NJH (2001) Intra-mitochondrial recombination: Is it why some mitochondrial genes sleep around?. Trends Ecol. Evol 16: 269-271. https://doi.org/10.1016/s0169-5347(01)02182-6
- Engel MS (1999) The taxonomy of Recent and fossil honey bees (Hymenoptera: Apidae: Apis). Journal of Hymenoptera Research 8: 165–196.
- Engel MS (2005) Family-group names for bees (Hymenoptera: Apoidea). Amer Mus Novitates 3476: 1–33. https://doi.org/10.1206/0003-0082(2005)476[0001:fnfbha]2.0.co;2
- Engel MS (2011a) Systematic melittology: where to from here?. Syst Entomol 36: 2–15. https://doi.org/10.1111/j.1365-3113.2010.00544.x
- Estoup AL, Solignac M, Harry M, Cornuet JM (1993) Characterization of (GT)n and (CT)n microsatellites in two insect species: Apis mellifera and Bombus terrestris. Nucleic Acids Research 21: 1427-1431. https://doi.org/10.1093/nar/21.6.1427
- Foltan P, Sheppard SK, Konvicka M, Symondson WOC (2005) The significance of facultative scavenging in generalist predator nutrition: detecting decayed prey in the guts of predators using PCR. Molecular Ecology 14: 4147-4158. https://doi.org/10.1111/j.1365-294x.2005.02732.x
- Franck P, Garnery L, Celebrano G, Solignac M, Cornuet JM (2000a) Hybrid origins of honey bees from Italy (Apis mellifera ligustica) and Sicily (A. m. sicula). Molecular Ecology 9: 907-921. https://doi.org/10.1046/j.1365-294x.2000.00945.x
- Franck P, Garnery L, Loiseau A (2001) Genetic diversity of the honeybee in Africa: microsatellite and mitochondrial data. Heredity 86: 420-430. https://doi.org/10.1046/j.1365-2540.2001.00842.x
- Franck P, Garnery L, Solignac M, Cornuet JM (2000b) Molecular confirmation of a fourth lineage in honey bees from Near East. Apidologie 31: 167-180. https://doi.org/10.1051/apido:2000114
- Gallai N, Salles JM, Settele J, Vaissie’re BE (2009) Economic valuation of the vulnerability of world agriculture confronted with pollinator decline. Ecol Econom 68: 810–821. https://doi.org/10.1016/j.ecolecon.2008.06.014
- Garnery L, Cornuet JM, Solignac M (1992) Evolutionary history of the honeybee Apis mellifera inferred from mitochondrial DNA analysis. Mol Ecology 1: 145–154. https://doi.org/10.1111/j.1365-294x.1992.tb00170.x
- Garnery L, Franck P, Baudry E, Vautrin D, et al. (1998) Genetic diversity of the west European honey bee (Apis mellifera and A. m. iberica). I. Mitochondrial DNA. Genetics Selection Evolution 30: 31-47. https://doi.org/10.1186/1297-9686-30-s1-s31
- Garnery L, Solignac M, Celebrano G, Cornuet JM (1993) A simple test using restricted PCR-amplified mitochondrial-DNA to study the genetic-structure of Apis mellifera L. Experientia 49: 1016-1021. https://doi.org/10.1007/bf02125651
- Gurney T, Elbel R, Ratnapradipa D, Brossard R (2000) Introduction to the molecular phylogeny of insects. Tested studies for laboratory teaching. Proceedings of the 21st Workshop/Conference of the Association for Biology Laboratory Education 21: 63-79.
- Haddad N, Meixner M, Fuchs S, Migdadi H, et al. (2009) Mitochondrial DNA support for genetic reserves of Apis mellifera syriaca in Jordan. Journal of Apicultural Research and Bee World 48: 19-22. https://doi.org/10.3896/ibra.1.48.1.05
- Haddada N, Adjlaneb N, Loucif-Ayadc W, Dashd A, et al. (2017) Mitochondrial genome of the North African Sahara Honeybee, Apis mellifera sahariensis (Hymenoptera: Apidae). Mitochondrial DNA Part B: Resources 2: 548–549. https://doi.org/10.1080/23802359.2017.1365647
- Hall HG (1990) Parental Analysis of Introgressive Hybridization Between African and European Honeybees Using Nuclear DNA RFLPs. Genetics 125: 611-621.
- Hall TA (1999) BioEdit: a user-friendly biological sequence alignment editor and analysis program for Windows 95/98/NT. Nucleic Acids Symp Ser. 41: 95-98.
- Hayashi M and Sota T (2010) Identification of Elmid larvae (Coleoptera: Elmidae) from Sanin District of Honshu, Japan, based on mitochondrial DNA sequences. Entomological Science 13: 417-424. https://doi.org/10.1111/j.1479-8298.2010.00404.x
- Hunt JG and Page ER (1992) Patterns of inheritance with RAPD molecular markers reveal novel types of polymorphism in the honey bee. Theoretical and Applied Genetics 85: 15-20. https://doi.org/10.1007/bf00223839
- Kajobe R (2006) Pollen foraging by Apis mellifera and stingless bees Meliponula bocandei and Meliponula nebulata in Bwindi Impenetrable National Park, Uganda. African Journal of Ecology 45: 265–274. https://doi.org/10.1111/j.1365-2028.2006.00701.x
- Klein AM, Vaissière BE, Cane JH, Steffan-Dewenter I, et al. (2007) Importance of pollinators in changing landscapes for world crops. Proceedings of the Royal Society B: Biological Sciences, 274: 303–313. https://doi.org/10.1098/rspb.2006.3721
- Kozmus P, Jevrosima S, Stanimirovic Z, Stojic V, et al. (2007) Analysis of mitochondrial DNA in honeybees (Apis mellifera) fr0m Serbia. Acta Veterinaria 57: 465-476. https://doi.org/10.2298/avb0706465k
- Magnus R and Szalanski AL (2010) Genetic evidence for honeybees (Apis mellifera L.) of Middle Eastern lineage in the United States. Sociobiology 55: 285.
- Melo GAR (1999) Phylogenetic relationships and classification of the major lineages of Apoidea (Hymenoptera), with emphasis on the crabronid wasps. Univ Kans Mus Nat Hist Sci Pap 14: 1–55. https://doi.org/10.5962/bhl.title.4053
- Michener CD (2007) The Bees of the World, second ed. John Hopkins University Press, Baltimore, Maryland (pp. xvi+[i]+953, +20 pls).
- Miguel I, Baylac M, Iriondo M, Manzano C, et al. (2011) Both geometric morphometric and microsatellite data consistently support the differentiation of the Apis mellifera M Evolutionary Branch. Apidologie 42: 150-161. https://doi.org/10.1051/apido/2010048
- Miguel I, Iriondo M, Garnery L, Sheppard WS, et al. (2007) Gene flow within the M evolutionary lineage of Apis mellifera: role of the Pyrenees, isolation by distance and post-glacial recolonization routes in the western Europe. Apidologie 38: 141-155. https://doi.org/10.1051/apido:2007007
- Mindell DP, Sorenson MD, Dimcheff DE (1998) Multiple independent origins of mitochondrial gene order in birds. Proc Natl Acad Sci USA 95: 10693-10697. https://doi.org/10.1073/pnas.95.18.10693
- Moritz RFA, Hawkins CF, Crozier RH, Mckinley AG (1986) A mitochondrial DNA polymorphism in honeybees (Apis mellifera L.). Experientia 42: 322-324. https://doi.org/10.1007/bf01942522
- Nei M and Kumar S (2000). Molecular Evolution and phylogenetics. Oxford University Press, New York.
- Oldroyd BP, Cornuet JM, Rowe D, Rinderer ET, et al. (1995) Racial admixture of Apis mellifera in Tasmania, Australia: similarities and differences with natural hybrid zones in Europe. Heredity 74: 315-325. https://doi.org/10.1038/hdy.1995.46
- Oldroyd BP and Wongsiri S (2006) Asian honeybees: biology, conservation and human interactions. Harvard University Press, Cambridge, MA, USA.
- Ozdil F, Fakhri B, Meydan H, Yildiz MA, et al. (2009) Mitochondrial DNA variation in the CoxI-CoxII intergenic region among Turkish and Iranian honey bees (Apis mellifera L.). Biochemical Genetics 47: 717-721. https://doi.org/10.1007/s10528-009-9269-8
- Papachristoforou Rortais A, Bouga M, Arnold G, Garnery L (2013) Genetic characterization of the Cyprian Honey bee (Apis mellifera cypria) based on microsatellites and mitochondrial DNA polymorphisms. Journal of Apicultural Sciences 57: 127-134. https://doi.org/10.2478/jas-2013-0023
- Pedersen BV (1996) On the phylogenetic position of the Danish strain of the black honeybee (the Laeso bee), Apis mellifera mellifera L. (Hymenoptera: Apidae) inferred from mitochondrial DNA sequences. Entomologica Scandinavica 27: 241-250. https://doi.org/10.1163/187631296x00070
- Prentice MA (1998) The comparative morphology and phylogeny of Apoid wasps (Hymenoptera: Apoidea). PhD thesis. Univ Calif Berkeley 1,439 pp.
- Ratcliffe NA, Mello CB, Garcia ES, Butt TM, Azambuja P (2011) Insect natural products and processes: New treatments for human disease. Insect Biochemistry and Molecular Biology 41(10): 747-769. https://doi.org/10.1016/j.ibmb.2011.05.007
- Rattanawannee A, Chanchao C, Wongsiri S (2010) Gender and Species Identification of Four Native Honey Bees (Apidae: Apis) in Thailand Based on Wing Morphometic Analysis. Annals of Entomological Society of America 103: 965-970. https://doi.org/10.1603/an10070
- Rortais A, Arnold G, Alburaki M, Legout H, et al. (2011) Review of the DraI COI-COII test for the conservation of the black honeybee (Apis mellifera mellifera). Conservation Genetic Resources 3: 383-391. https://doi.org/10.1007/s12686-010-9351-x
- Rukhsana K, Akhilesh VP, Sebastian CD (2014) Deciphering the molecular phylogenetics of the Asian honey bee, Apis cerana and inferring the phylogeographical relationships using DNA barcoding. Journal of Entomology and Zoology Studies 2: 218-220.
- Ruttner F (1988) Biogeography and taxonomy of honey bees. Springer, Berlin, Germany, 13-19. https://doi.org/10.1007/978-3-642-72649-1_2
- Ruttner F (1992) Naturgeschichte der Honigbienen. Ehrenwirth Verlag. Münich. Germany. 357 pp.
- Schiff NM and Sheppard W (1995) Genetic Analysis of Commercial Honey Bees (Hymenoptera: Apidae) from the Southeastern United States. Economic Entomology 88: 1216-1220. https://doi.org/10.1093/jee/88.5.1216
- Shao R, Dowton M, Murrell A, Barker SC (2003) Rates of gene rearrangement and nucleotide substitution are correlated in the mitochondrial genomes of insects. Mol. Biol. Evol. 20: 1612-1619. https://doi.org/10.1093/molbev/msg176
- Sheppard W and Smith D (2000) Identification of African derived bees in the Americas: a survey of methods. Annals of the Entomological Society of America 93: 159-176. https://doi.org/10.1603/0013-8746(2000)093[0159:ioadbi]2.0.co;2
- Sheppard WS, Arias MC, Grech A, Meixner MD (1997) Apis mellifera ruttneri, a new honey bee subspecies from Malta. Apidologie 28: 287–293. https://doi.org/10.1051/apido:19970505
- Sheppard WS and Meixner MD (2003) Apis mellifera pomonella, a new honeybee subspecies from Central Asia. Apidologie 34: 367–375. https://doi.org/10.1051/apido:2003037
- Silvestre D and Arias MC (2006) Mitochondrial tRNA gene translocations in highly eusocial bees. Genet Mol Biol 29: 572-575. https://doi.org/10.1590/s1415-47572006000300030
- Silvestre D, Francisco FO, Weinlich R, Arias MC (2002) A scientific note on mtDNA gene order rearrangements among highly eusocial bees. Apidologie 33: 355-356. https://doi.org/10.1051/apido:2002021
- Smith D (1991) Mitochondrial DNA and honey bee biogeography. In: Smith D.R. (Ed.) Diversity of the Genus Apis. Westview Press. Oxford and IBH Publishing Co Oxford: 131-176.
- Smith DR, Hagen RH (1997) The biogeography of Apis cerana as revealed by mitochondrial DNA sequence data. Journal of the Kansas Entomological Society 69: 294-310.
- Smith DR, Taylor OR, Brown WM (1989) Neotropical Africanized honeybees have African mitochondrial DNA. Nature 339: 213-215. https://doi.org/10.1038/339213a0
- Smith MA, Woodley NE, Janzen DH, Hallwachs W, et al. (2006) DNA barcodes reveal cryptic host specificity within the presumed polyphagous members of a genus of parasitoid flies (Diptera: Tachinidae). Proceedings of National Academy of Sciences USA 103: 3657-3662. https://doi.org/10.1073/pnas.0511318103
- Solignac M, Vautrin D, Baudry E, Mougel F, Loiseau A, Cornuet JM (2004) A microsatellite-based linkage map of the honeybee, Apis mellifera L. Genetics 167: 253-262. https://doi.org/10.1534/genetics.167.1.253
- Solorzano CD, Szalanski AL, Kence M, Mckern JA, et al. (2009) Phylogeography and population genetics of honeybees (Apis mellifera) from Turkey based on COI-COII sequence data. Sociobiology 53: 237-246.
- Soroker V, Hetzroni A, Yacobson B, Voet H, et al. (2009) Colony losses in Israel: incidence of viral infection and beehive populations. In: Proceedings of the 41st Apimondia Congress, Mointpellier, France. p. 38.
- Szalanski AL and Magnus RM (2010) Mitochondrial DNA characterization of Africanized honeybee (Apis mellifera L.) populations from the USA. Journal of Apicultural Research 49: 177-185. https://doi.org/10.3896/ibra.1.49.2.06
- Tamura K, Stecher G, Peterson D, Filipski A, et al. (2013) MEGA6 molecular evolutionary genetics analysis version 60. Mol Biol Evol. 30: 2725–2729. https://doi.org/10.1093/molbev/mst197
- Tarès S, Cornuet JM, Abad P (1993) Characterization of an Unusually Conserved Alu I Highly Reiterated DNA Sequence Family From the Honeybee Apis mellifera. Genetics 134: 1195-1204.
- Techer MA, Clémencet J, Turpin P, Volbert N, et al. (2015) Genetic characterization of the honeybee (Apis mellifera) population of Rodrigues Island, based on microsatellite and mitochondrial DNA. Apidologie 46: 445-454. https://doi.org/10.1007/s13592-014-0335-9
- The Honey Bee Genome Sequencing Consortium (HGSC) (2006) Insights into social insects from the genome of the honeybee Apis mellifera. Nature 443: 931-949. https://doi.org/10.1038/nature05260
- Thompson JD, Gibson TJ, Plewniak F, Jeanmougin F, et al. (1997) The CLUSTAL-X windows interface: flexible strategies for multiple sequence alignment aided by quality analysis tools. Nucleic Acids Res 25: 4876–4882. https://doi.org/10.1093/nar/25.24.4876
- Van Engelsdorp D, Evans JD, Saegerman C, Mullin C, et al. (2009) Colony collapse disorder: a descriptive study. PloS ONE 4: e6481. https://doi.org/10.1371/journal.pone.0006481
- Van Engelsdorp D and Meixner MD (2010) A historical review of managed honeybee populations in Europe and the United States and the factors that may affect them. J Invertebr Pathol 103: 80–95. https://doi.org/10.1016/j.jip.2009.06.011
- Whitfield CW, Behura SK, Berlocher SH, Clark AG, et al. (2006). Thrice out of Africa: ancient and recent expansions of the honey bee, Apis mellifera Science 314: 642-645. https://doi.org/10.1126/science.1132772
- Williams I (2002) Insect Pollination and Crop Production: A European Perspective. In P. Kevan & V. L. Imperatriz Fonseca (Eds.), Pollinating Bees - The Conservation Link Between Agriculture and Nature Ministry of Environment / Brasília, pp. 59-65.
Keywords:
Download:
Full PDF- Share This