Genotypes within the prolactin and growth hormone insulin-like growth factor-I pathways associated with milk production in heat stressed Holstein cattle: Genotypes and milk yield in heat stressed Holstein cows
Received: October 12, 2017
Accepted: October 12, 2017
Published: October 31, 2017
Genet.Mol.Res. 16(4): gmr16039821
DOI: 10.4238/gmr16039821
Abstract
The association of single nucleotide polymorphisms (SNPs) within genes of the prolactin (PRL), growth hormone (GH) and insulin-like growth factor-I (IGF1) pathways with 305-day milk yield (MY305) were evaluated in heat-stressed Holstein cows. Ambient temperature and relative humidity were used to calculate a temperature-humidity index (THI), which revealed that heat stress conditions existed in the Yaqui Valley, Sonora, México from May through October of 2011 and 2012. Cows (n = 573) were genotyped for 179 tag SNPs within 43 candidate genes in the PRL and GH/IGF1 pathways. Seven SNPs within 7 genes (AVPR1A, Furin, IGFBP5, IGFBP6, PMCH, PRLR and STAT5B) were found to be associated with MY305 (P<0.05); therefore, their effects were used to estimate a molecular breeding value (MBV). The correlation between the MBV and MY305 was 0.21 (P<0.001) and the adjusted coefficient of determination (R2) was 4%, whereas correlation between MBV and the estimated breeding value for MY305 was 0.25 (P<0.001) with a R2 of 6%. Heritability estimate for MY305 was 0.41 ± 0.12. The MBV estimated was positive, but weakly associated with MY305. The small amount of additive and phenotypic variation explained by the MBV was most likely due to the few number of SNPs and the complexity of the trait, particularly under extreme weather conditions. In conclusion, SNPs within the PRL and GH/IGF1 pathways were associated with MY305; however, the MBV estimated with these SNPs was not yet suitable to use in genetic selection procedures due to the small amount of variation explained for MY305.
References
Yacht Charter in Balkans Blog | Casino Sites in Bangladesh Yacht Charter in Australia Blog | Mortgage info in New York Yacht Charter in China Blog | Best Betting Sites in Nigeria Yacht Charter in Spain Blog | Best Betting Sites in United Arab Emirates Yacht Charter in France Blog | Places to Visit in China Yacht Charter in Greece Blog | Best Betting Sites in Indonesia Yacht Charter in Croatia Blog - Find Lawyer in Connecticut Yacht Charter in Italy Blog - Find Lawyer in Maryland Yacht Charter in Japan Blog | Info Blockchain Technologies Yacht Charter in Philippines Blog | Obgyn JournalsIntroduction
One of the major challenges for dairy cattle in tropical, subtropical and semi-arid regions is heat stress. Environmental conditions such as high ambient temperatures, high relative humidity and prolonged periods of solar radiation, compromises the ability of dairy cows to dissipate heat. This condition causes behavioral changes such as increased water consumption and decreased feed intake, which in turn, results in losses in reproductive efficiency and milk production (Accorsi et al., 2002; West, 2003; Bernabucci et al., 2010). Persistent exposure to heat stress leads cows to an altered physiological state known as acclimation, which is a process largely controlled by the endocrine system that involves changes in plasma concentration of several hormones and somatomedins like prolactin (PRL), growth hormone (GH), insulin-like growth factor-I (IGF1), thyroid hormones, glucocorticoids, and mineral corticoids (Collier et al., 2008; Bernabucci et al., 2010). The PRL hormone plays a key role in lactogenesis in ruminants and some reports indicate that the synthesis and secretion of PRL by the anterior pituitary gland is sensitive to changes in ambient temperature, specifically higher levels of this hormone are observed during summer due to its influence in regulation of body fluids and its relationship with seasonal hair growth (Alamer, 2011). In addition to PRL, other important biological molecules altered during heat stress are GH and IGF1. These hormones are members of the somatotropic axis which regulate metabolism and physiology of mammalian growth, however, they are affected by the negative energy balance derived from the reduction of feed intake in fresh cows that is exacerbated by heat stress (De Rensis and Scaramuzzi, 2003; Bonakdar et al., 2010).
The genetic development of heat tolerant dairy cattle is complex; however, with advancements of molecular and genomic technologies it is posible to identify DNA markers (SNPs) within candidate genes that underlie quantitative trait loci (QTL) that contribute to the favorable phenotypic variations of animals that have fitness in heat stress conditions (Bonakdar et al., 2010; Shaji et al., 2015). The hypothesis of this study, was that a molecular breeding value (MBV) constructed with DNA markers within genes of the PRL and GH/IGF1 pathways may have the potential to predict 305-day milk yield in heat stressed Holstein dairy cows. To test this hypothesis, 179 tag SNP were identified in 43 genes within these pathways. The objectives of the study were to estimate an MBV using these SNPs and evaluate their association with 305-day milk yield in heat-stressed Holstein cows. Another objective was to estimate the amount of additive and phenotypic variability explained by the MBV for 305-day milk yield in heat-stressed lactating Holstein cows on 3 dairy farms in Sonora, México.
Material and Methods
Data
Data were collected from 573 Holstein dairy cows. Observations were collected during 2011 and 2012 from three dairy farms located in Blocks 910 and 1114 of the Yaqui Valley, Sonora, México (27°21'N 109°54W). Each farm contributed the following number of animals for the study: farm 1 (n = 252), farm 2 (n = 89), and farm 3 (n = 232). Cows had body condition scores ranging from 2.5 to 3.5 (1 = very thin, 5 = very fat) (Kadarmideen and Wegmann, 2003). Test-day milk yield records were used to calculate an adjusted 305-day milk yield (MY305) per cow in kg. The MY305 was calculated by multiplying milk production levels by an adjustment factor that depended on days in milk and the age of the cow. These factors were obtained from the Dairy Herd Improvement Association (McDaniel et al., 1965; Norman et al., 1973).
Management and health status
As part of a routine management of all farms during summer, cows were subjected to showers with cooling systems as was described for the control group in the study of Leyva-Corona et al. (2016) to minimize the detrimental effects of heat stress. Cooling systems had 16 sprinklers (with an approximate water distribution of 15L per cow/cycle) and three ~ 367 watts electrical fans located 2.73 meters above the floor. Cows were maintained in shaded holding pens with free access to water and an average of 8.5 m2 of shade per cow. In farms 1 and 3, each holding pen contained approximately 50 cows, meanwhile in farm 2, the holding pens contained 25 to 30 animals. Cows were fed twice a day with a ration consisting of 75% alfalfa hay and 25% corn silage that supplied their nutritional requirements according to guidelines established by the NRC (2001), for lactating Holstein cows with an average weight of 650 kg and producing approximately 30 kg/d of milk with average composition of 3.5% fat and 3.2% protein.
Health records were obtained from the cows of the three dairies and were compiled into one variable known as health status that was recorded categorically and coded as 0 for no disease diagnosis and 1 for any disease diagnosis. Metritis diagnosis was performed with a portable ultrasound system (SonoSite, Inc., Bothell, WA) at day 40 postpartum, while for the diagnosis of subclinical mastitis, the qualitative California mastitis test (Laboratorios Sanfer, S.A. de C.V., Obregón, México) was used. Summary statistics (Table 1) were calculated for each health status group using PROC MEANS of SAS (version 9.4; SAS Institute Inc., USA).
| Health | Variables | N | Mean | SD | Minimum | Maximum |
|---|---|---|---|---|---|---|
| 0 | MY3051, kg | 545 | 6412.22 | 1392.10 | 2863.8 | 10787 |
| Years of age | 544 | 5.34 | 1.94 | 2 | 13 | |
| Number of lactation | 544 | 3.14 | 1.85 | 1 | 11 | |
| DIM2 | 545 | 307 | 47 | 116 | 471 | |
| 1 | MY3051, kg | 28 | 5514.45 | 1580.72 | 2314.72 | 8029.77 |
| Years of age | 28 | 4.99 | 1.75 | 2 | 10 | |
| Number of lactation | 28 | 2.82 | 1.59 | 1 | 8 | |
| DIM2 | 28 | 308 | 75 | 122 | 441 |
1MY305 = 305-day milk yield. 2DIM = Days in milk
Table 1: Summary statistics per health status group (0 is for no disease diagnosis, 1 for any disease diagnosis) in lactating Holstein cows in Sonora, MX.
Temperature and humidity index (THI)
The THI was calculated and provided by the Instituto Tecnológico de Sonora for the years 2011 and 2012. The climatic records were obtained thru Sonora, México Agro-climatic Station Network, available through www.agroson.org México. This calculation was based on the equation:
THI = 0.81(T°C) + RH /100(T°C -14.4) + 46.4
where THI represented the temperature and humidity index, T°C was the temperature in degrees Celsius and RH was the percentage of relative humidity (Hahn, 1999). Index values were calculated for each hour of each day (24 times), and then, were used to calculate monthly averages to plot the trajectory of the THI thru the years of study.
SNP discovery and genotypes
Forty-three candidate genes (see Table S1) within the PRL and GH/IGF1 pathways were studied. These genes were selected based on their physiological function, involvement in milk production and documented heat stress responsiveness (Accorsi et al., 2002; Etherton, 2003). For DNA extraction, blood samples (3 ml) were obtained by caudal venipuncture of each cow using sterile syringes. This sample was spotted on nucleic acid cards (GeneSeek, Inc., Lincon, NE). Genotyping was then completed using several multiplex SNP assays and the Sequenom Mass Array platform (GeneSeek, Inc., Lincon, NE). Linkage disequilibrium regions between the genotyped polymorphisms were analyzed using HaploView software (Barrett et al., 2005). A range from 2 to 50 SNPs were found within each gene and yielded a panel of 179 tag SNPs (see Table S2).
SNP effects
The associative analysis between genotypes and phenotypes for MY305 were performed using the mixed procedure of SAS. The statistical model was:
y = Xb + Za + e
where y corresponded to the 305-day milk yield observation of each cow, b was the vector of unknown fixed effects, and a was the vector of unknown random sire effects. Fixed effects included SNP genotype (0, 1 or 2) modelled as a categorical variable, farm, lactation number, health status and calving month; days in milk were included as a linear covariate. X and Z were known incidence matrices relating observations in y to both fixed and random effects, and e was the vector of unknown residual errors. Associations between each SNP and 305-day milk yield were reported based on their significance (P < 0.05). Each single SNP effect was estimated using two distinct approaches. In the first approach, the SNP genotype was included as a covariate to determine the allele substitution effect. In the second approach, the SNP genotype was included in the model as a categorical variable and orthogonal contrasts were used to estimate additive effects (Cochran et al., 2013). The false discovery rate (FDR) or Q-value was calculated to control for false positives using PROC MULTTEST (Benjamini and Hochberg, 1995).
Molecular breeding value (MBV)
The MBV was calculated only for the cows that had a genotyping rate of 100% for all the SNPs that showed a significant independent association (P < 0.05) with MY305 (n = 538 cows). The MBV estimation involved the summation of the additive genotype effect of each locus and was performed using the Animal Breeder Tool Kit (Golden et al., 1992). Pearson’s correlation between 305-day milk yield and the MBV, as well as the EBV for MY305 and the MBV were calculated using PROC CORR in SAS (version 9.4; SAS Institute Inc., USA). A regression-prediction analysis using PROC MIXED was used to estimate a full and a reduced model in order to calculate the amount of phenotypic variation explained by the MBV. The full model was:
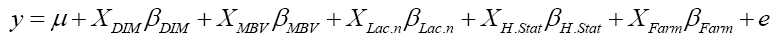
where y was the dependent variable of 305-day milk yield, μ was the population mean, XDIM was the linear covariate for days in milk, βDIM was the slope for the variable days in milk, XMBV was the linear covariate for MBV, βMBV was the slope for the variable MBV,XLac.nβLac.n, XH.StarβH.Star, XFarmβFarm were the incidence matrixes for the categorical variables number of lactations, health status and farm with vectors for fixed effects respectively and e was the vector of residual effect or error term.
The reduced model constructed only with the dependent variable and the MBV was:
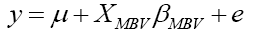
where y was defined as the dependent variable of 305-day milk yield, μ was the population mean, XMBV was the linear covariate for MBV and βMBV was the slope for the variable MBV, and e was the vector of residual effect or error term.
Estimated Breeding Value and Heritability for 305-day milk yield
A mixed model was implemented using ASREML 3.0 (Gilmour et al., 2009) in order to estimate breeding values (EBV) and the heritability of 305-day milk yield. The general model equation was:
y = Xb + Za + e
where y was the vector of 305-day milk yield, b was the vector of fixed effects, and a was the vector of random effects which included the individual cow. Fixed effects included farm, lactation number, health status and calving month. The variable days in milk were included as linear covariate. The X and Z were known incidence matrices relating records in y to both fixed and animal random effects in b and a , and e was the vector of random residual effects. The pedigree file included a total of 971 individuals. In order to estimate the amount of additive variance explained by the MBV, a final regression-prediction analysis using PROC MIXED was used. The model was:
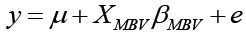
where y was the EBV for 305-day milk yield obtained from the previous model, μ was the population mean, XMBV was the covariate for MBV and βMBV was the slope for the variable MBV, and e was the vector of residual effect or error term. It is important to mention that this model was implemented using only cows that had a valid observation for the MBV.
Results
Temperature humidity index (THI)
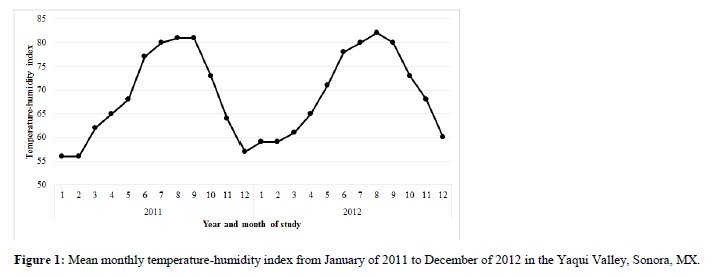
Estimated THI values revealed that cows in this study were exposed to heat stress conditions from approximately May until October during both years of the study (Figure 1), varying from light (72-79 units) to moderate stress (80-89 units) according to the classification described by Armstrong (1994).
SNP effects
Seven SNPs within 7 genes were associated (P < 0.05) with 305-day milk yield. These genes were the Arginine vasopressin receptor 1A (AVPR1A), Furin, Insulin-like growth factor binding proteins 5 and 6 (IGFBP5, IGFBP6), Pro-melanin concentrating hormone (PMCH), Prolactin receptor (PRLR) and the Signal Transducer and Activator of Transcription 5B (STAT5B). Table 2 shows the significance level (P-value), chromosomal location and additive effect for each SNP.
| Genes | Chr1 | Location (Mb) | SNP | P | FDR2 | Alleles | Additive effect (Kg) |
|---|---|---|---|---|---|---|---|
| IGFBP5 | 2 | 10.5 | rs208989155 | 0.02 | 0.03 | A / G | 494.97 |
| AVPR1A | 5 | 50.5 | rs210011420 | 0.02* | 0.03 | T / C | 286.75 |
| IGFBP6 | 5 | 27.0 | rs211039223 | 0.02* | 0.03 | C / T | 332.42 |
| PMCH | 5 | 66.3 | rs135033882 | 0.01* | 0.03 | A / T | 37.92 |
| STAT5B | 19 | 42.9 | rs384930401 | 0.01 | 0.03 | G / A | 373.34 |
| PRLR | 20 | 39.1 | rs135164815 | <0.01* | 0.03 | A / G | 222.93 |
| FURIN | 21 | 22.2 | rs381099643 | 0.03* | 0.03 | G / A | 253.11 |
Favorable alleles are bolded.1Chr = chromosome. 2FDR = false discovery rate.
Table 2: SNPs within the PRL and GH-IGF1 pathways associated with 305-day milk yield in heat stressed lactating Holstein cows in Sonora, MX.
MBV
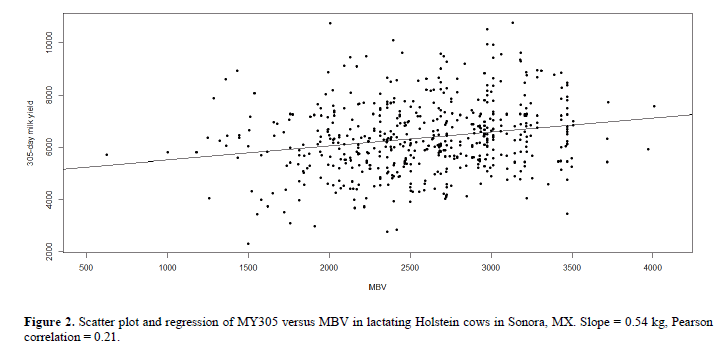
The molecular breeding value was calculated for 538 cows that were genotyped for all the SNPs that were found to have a significant association with MY305 (Figure 2). The resulting MBV had an average value of 2575.92 and ranged from 623.45 to 4008.88. Within the full model, all variables included were statistically significant for predicting MY305, this model had an adjusted coefficient of determination (R2) of 47.2%. The reduced model constructed with our variable of interest (MBV) as an independent variable had an adjusted R2 of 4%, this value represents the percentage of the phenotypic variance explained by the MBV.
Genetic parameters
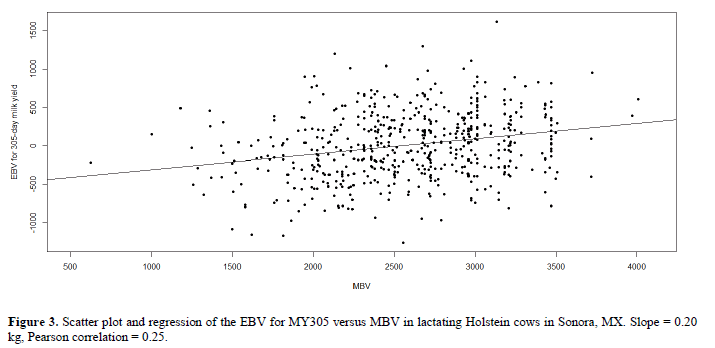
Additive genetic and phenotypic variances as well as heritability were estimated for 305-day milk yield. Our study revealed a heritability estimate of 0.41 with a standard error of 0.12 for 305-day milk yield (Table 3). The correlation between the MBV and the MY305 EBV was 0.25 (P < 0.0001) while regression of the MY305 EBV on the MBV is shown in Figure 3. The percentage of the additive variance explained by the MBV (represented by the adjusted R2) was 6%.
| Variables | MY3051 (kg2) | SE (kg) |
|---|---|---|
| σ2a | 439980 | 140380 |
| σ2e | 621100 | 124430 |
| σ2p | 1061080 | 66803 |
| h2 | 0.41 | 0.12 |
Table 3: Genetic parameters for 305-day milk yield in lactating Holstein cows in Sonora, MX. without using the MBV as a fixed effect.
Discussion
Based on the trajectory of the THI thru the years of study, it should be noted that the environmental conditions during the summer in the Yaqui Valley exceeded 72 THI units, which is considered the threshold to initiate heat stress in dairy cattle (Armstrong, 1994). These results agree with other studies conducted in this geographical region of México that described that the THI can be moderate and reach around 85 units during summer (Rivera-Acuña et al., 2015). To help mitigate the effect indicated by the THI on dairy cattle, cooling systems are used in northwest México (Leyva-Corona et al., 2016); however, it has also been reported that even with the use of cooling systems, milk yield decreases by 10 to 15% when ambient temperature causes heat stress (Dunshea et al., 2013). These previous findings have shown the increased need to find new strategies that could allow the improvement of milk production in these climate zones; concomitantly, Collier et al. (2008) described the important role of genetics in the bovine heat stress response. In parallel with this review of heat-stress responsive endocrine pathways, we previously reported suppressed serum concentrations of PRL in Holstein cows being milked in the summer in the Yaqui Valley (Leyva-Corona et al., 2016). Serum levels of IGF1 were also influenced by cooling strategies on these dairy farms. These results were from an intensive endocrine study of heat-stressed Holstein cows; however, for the much larger scale genotype to phenotype association study of the current effort, it was unfeasible to obtain intensive endocrine data for 573 lactating cows.
Our findings revealed a SNP (rs210011420) located at the AVPR1A gene associated with 305-day milk yield in heat stressed Holstein cows. To our knowledge, no previous research regarding this genotype to phenotype association exists. The AVPR1A gene is located on bovine chromosome 5 (BTA5) and is involved in regulation of systemic arterial pressure. The previous could be the reason of its involvement in the physiological response to heat stress in cattle, because when cows are subjected to heat stress, the blood flow is shifted to peripheral tissues for cooling (West, 2003). Moreover, the protein AVPR1A is a receptor for the arginine vasopressin hormone (AVP), which is a diuretic protein involved in the secretion and ejection of milk during lactation (Nussey et al., 1987), which could explain its association with 305-day milk yield.
Other genetic variants that were found to be associated with 305-day milk yield were rs208989155 and rs211039223, located within the genes IGFBP5 and IGFBP6, respectively. Their association may be explained by the role of these binding proteins in the somatotropic axis, since the activity of IGF1 (i.e., free vs bound form in circulation) in milk production and cell proliferation during lactation is regulated by the IGFBP protein family (Accorsi et al., 2002). Specifically, it has been reported that IGFBP5 is an important regulator of apoptosis in the mammary gland and that IGFBP6 shows a decrease in its expression during lactation (Accorsi et al., 2002; Fenwick et al., 2008).
This research reported evidence of the association between the SNP (rs381099643) in the Furin gene and 305-day milk yield. The association may be explained by Furin involvement in the post-translational processing of growth hormone release hormone (GHRH) and, indirectly, in the synthesis and secretion of GH (Posner et al., 2004), which could affect nutrient mobilization and cell proliferation during lactation. Furin is involved in the activation of precursor proteins through the cleavage of a single or paired basic amino acid residue (Khatib and Sfaxi, 2012; Maruotti et al., 2012). Previous research investigated Furin in lactating cows (Cánovas et al., 2010), but an association with milk production traits was not reported.
Previous research has not identified SNP within the PMCH gene to be associated with milk production traits. In our study, one SNP (rs135033882) was found to be associated with 305-day milk yield. This association may be explained by the effect of this gene on energy status, which could lead to changes in energy balance in heat-stressed cows. The PMCH gene is involved in regulation of energy homeostasis and could be a defense mechanism against energy deficiency (Beerda et al., 2008). It should be noted that energy balance decreases rapidly within the first 100 days in milk in high-yield lactating Holstein cows (Hüttmann et al., 2009).
Another polymorphism associated with 305-day milk yield was rs384930401, which is located in the STAT5B gene. Similar results were provided by He et al. (2011), who indicated that a SNP within the STAT5B gene was significantly associated with the EBV for milk production in Chinese Holstein cattle.
An important candidate gene identified as being associated with 305-day milk yield was PRLR. The relevance of PRLR is due to its role in milk production and heat stress response (Collier et al., 2008, Lü et al., 2010). The SNP rs135164815 within the PRLR gene that was associated with milk yield in our research was located in exon 2, position 39.1 Mb on BTA20. This result is similar to the reported by Zhang et al. (2008), who reported that polymorphisms in exons 3 and 7 of the PRLR gene were associated with milk yield in Holstein cattle. In contrast, other studies found a PRLR mutation in exon 10 that introduces a premature stop codon and is considered a candidate for slick hair coat genotype and heat tolerance in Senepol cattle. Specifically, this SNP (39.1 Mb) in the PRLR gene was a single-base deletion in exon 10 that introduced a premature stop codon at the protein position 462 (Leu) yielding a loss of 120 C-terminal amino acids from the long isoform of the receptor (Littlejohn et al., 2014).
Previous research identified a “phenotype characterized by development of a very short, sleek hair coat that is inherited as if controlled by a single dominant gene”, the slick gene (Olson et al., 2003). In that study, it was reported that Holstein dairy cattle with the slick haplotype exhibit higher milk yields than non-slick contemporaries. This is of particular interest for dairy farming, where most genetic selection has occurred in heat-intolerant Bos taurus breeds (Littlejohn et al., 2014). Moreover, PRLR is found at the same locus, BTA20, as other DNA markers used to map the slick gene in Bos taurus cattle. Within this region, two genes, PRLR (38.0 Mb) and sperm flagellar 2 (SPEF2) (38.4 Mb) were found. The SPEF2 and PRLR genes are also involved in reproduction and milk production. Additionally, values of integrated haplotype scores indicated that the region between PRLR and SPEF2 is a target of recent selection (Huson et al., 2014).
Our study revealed a heritability estimate of 0.41 ± 0.12 for 305-day milk yield (Table 3). Other studies reported heritabilities ranging from 0.29 to 0.37 for milk yield (Cohen-Zinder et al., 2005, Raven et al., 2013). Calculation of a moderate heritability confirms the important contribution of genetics to milk yield in Holstein cows. The MBV estimated from SNPs within the PRL and GH/IGF1 pathways genes was weakly associated with 305-day milk yield and with the EBV for 305-day milk yield in Holstein cows from Sonora, México. Although an alternative MBV was also calculated using all available SNPs, this estimate was not adequate, perhaps due to the lack of relationship between most of the variants with MY305. The predictive ability of the MBV for the phenotypic and the additive variances was extremely low, probably due to the few number of SNPs included in its estimation and the complexity of the trait, specially under heat stress conditions.
Conclusion
In conclusion, we found association between seven SNPs within the PRL and GH/IGF1 pathways and 305-day milk yield, therefore, the result of this study support our hypothesis that a molecular breeding value (MBV) constructed with DNA markers within the PRL and GH/IGF1 pathways may have the potential to predict 305-day milk yield in heat stressed Holstein dairy cows; however, due to the small amount of additive and phenotypic variability explained by the MBV, its implementation is not yet feasible in genetic selection procedures.
Acknowledgments
The authors thank to the dairy farmers of the Yaqui Valley for they valuable contribution and access to their herds, the Instituto Tecnológico de Sonora in Mexico for the data used, Colorado Ag Experimental Station and the department of Animal Science of Colorado State University for providing the tools to analyze the data, UC MEXUS program for funding this research project and the Fulbright program of scholarships for funding A. I. Hernández-Cordero.
About the Authors
Corresponding Author
M. G. Thomas
Department of Animal Sciences, Colorado State University, Fort Collins, CO, USA
- Email:
- milt.thomas@colostate.edu
References
- Accorsi PA, Pacioni B, Pezzi C, Forni M, et. al. (2002). Role of prolactin, growth hormone and insulin-like growth factor 1 in mammary gland involution in the dairy cow. J. Dairy Sci. 85: 507–513. https://doi.org/10.3168/jds.s0022-0302(02)74102-7
- Alamer M (2011). The role of prolactin in thermoregulation and water balance during heat stress in domestic ruminants. Asian J. Anim. Vet. Adv. 6: 1153-1169.https://doi.org/10.3923/ajava.2011.1153.1169
- Armstrong D (1994). Heat stress interaction with shade and cooling. J. Dairy Sci. 77: 2044-2050. https://doi.org/10.3168/jds.s0022-0302(94)77149-6
- Barrett JC, Fry B, Maller J and Daly MJ. (2005). Haploview: analysis and visualization of LD and haplotype maps. Bioinformatics. 21: 263- https://doi.org/10.1093/bioinformatics/bth457
- Beerda B, Wyszynska-Koko J, te Pas MFW, de Wit AAC, et al. (2008). Expression profiles of genes regulating dairy cow fertility: recent findings, ongoing activities and future possibilities. Animal. 2: 1158-1167. https://doi.org/10.1017/s1751731108002371
- Benjamini Y and Hochberg Y (1995). Controlling the false discovery rate: a practical and powerful approach to multiple testing. J. R. Stat. Soc. Series B Stat. Methodol.: 57: 289-300.
- Bernabucci U, Lacetera N, Baumgard LH, Rhoads RP, et al. (2010). Metabolic and hormonal acclimation to heat stress in domesticated ruminants. Animal. 4: 1167-1183. https://doi.org/10.1017/s175173111000090x
- Bonakdar E, Rahmani HR, Edriss MA and Tabatabaei BS (2010). IGF-I gene polymorphism, but not its blood concentration, is associated with milk fat and protein in Holstein dairy cows. Genet. Mol. Res. 9: 1726-1734. https://doi.org/10.4238/vol9-3gmr874
- Cánovas A, Rincon G, Islas-Trejo A, Wickramasinghe S, et al. (2010). SNP discovery in the bovine milk transcriptome using RNA-Seq technology. Mamm. Genome. 21: 592-598. https://doi.org/10.4238/vol9-3gmr874
- Cochran SD, Cole JB, Null DJ and Hansen PJ (2013). Discovery of single nucleotide polymorphisms in candidate genes associated with fertility and production traits in Holstein cattle. BMC Genet. 14: 49. https://doi.org/10.1186/1471-2156-14-49
- Cohen-Zinder M, Seroussi E, Larkin DM, Loor JJ, et al. (2005). Identification of a missense mutation in the bovine ABCG2 gene with a major effect on the QTL on chromosome 6 affecting milk yield and composition in Holstein cattle. Genome Res. 15: 936-44. https://doi.org/10.1101/gr.3806705
- Collier RJ, Collier JL, Rhoads RP and Baumgard LH (2008). Invited Review: Genes Involved in the Bovine Heat Stress Response. J. Dairy Sci. 91: 445-454.https://doi.org/10.3168/jds.2007-0540
- De Rensis F and Scaramuzzi RJ (2003). Heat stress and seasonal effects on reproduction in the dairy cow--a review. Theriogenology 60: 1139-51. https://doi.org/10.1016/s0093-691x(03)00126-2
- Dunshea FR, Leury BJ, Fahri F, DiGiacomo K, et al. (2013). Amelioration of thermal stress impacts in dairy cows. Anim. Prod. Sci. 53: 965-975. https://doi.org/10.1071/an12384
- Etherton T (2003). Somatotropic function: the somatomedin hypothesis revisited. J. Anim. Sci. 82(13_suppl): E239-244.
- Fenwick MA, Llewellyn S, Fitzpatrick R, Kenny DA, et al. (2008). Negative energy balance in dairy cows is associated with specific changes in IGF-binding protein expression in the oviduct. Reproduction. 135: 63-75. https://doi.org/10.1530/rep-07-0243
- Gilmour AR, Gogel BJ, Cullis BR, Thompson R, et al. (2009). ASReml user guide release 3.0. VSN International Ltd, Hemel Hempstead, UK.
- Golden, B. L., W. M. Snelling, and C. H. Mallinckrodt. 1992. Animal breeder’s toolkit user’s guide and reference manual. Agric. Exp. Stn.Tech. Bull. LTB92-2. Colorado State University, Fort Collins, CO.
- Hahn, G. L. 1999. Dynamic responses of cattle to thermal heat loads. J. Anim. Sci. 79 (Suppl. 2), 10-20. https://doi.org/10.2527/1997.77suppl_210x
- He Y, Chu Q, Ma P, Wang Y, et al. (2011). Association of bovine CD4 and STAT5b single nucleotide polymorphisms with somatic cell scores and milk production traits in Chinese Holsteins. J. Dairy Res. 78:242-249. https://doi.org/10.1017/s0022029911000148
- Huson HJ, Kim ES, Godfrey RW, Olson TA, et al. (2014). Genome-wide association study and ancestral origins of the slick-hair coat in tropically adapted cattle. Front. Genet. 5: 101. https://doi.org/10.3389/fgene.2014.00101
- Hüttmann H, Stamer E, Junge W, Thaller G, et al. (2009). Analysis of feed intake and energy balance of high-yielding first lactating Holstein cows with fixed and random regression models. Animal 3: 181-188. https://doi.org/10.1017/s175173110800325x
- Kadarmideen HN, Wegmann S (2003). Genetic parameters for body condition score and its relationship with type and production traits in Swiss Holsteins. J. Dairy Sci. 86: 3685:3693. https://doi.org/10.3168/jds.s0022-0302(03)73974-5
- Khatib AM, Sfaxi F (2012). FURIN (furin (paired basic amino acid cleaving enzyme). Atlas Genet. Cytogenet. Oncol. Haematol. 16: 639-643.
- Leyva-Corona JC, Thomas MG, Rincón G, Medrano JF, et al. (2016). Enfriamiento al inicio de verano para mitigar el estrés por calor en vacas Holstein del noroeste de México. Rev. Mex. Cienc. Pecu. 7: 415-429. https://doi.org/10.22319/rmcp.v7i4.4274
- Littlejohn MD, Henty KM, Tiplady K, Johnson T, et al. (2014). Functionally reciprocal mutations of the prolactin signalling pathway define hairy and slick cattle. Nat. Comm. 5: 5861. https://doi.org/10.1038/ncomms6861
- Lü A, Hu X, Chen H, Jiang J, et al. (2010). Single nucleotide polymorphisms in bovine PRL gene and their associations with milk production traits in Chinese Holsteins. Mol. Biol. Rep. 37: 547-551. https://doi.org/10.1007/s11033-009-9762-5
- Maruotti J, Muñoz M, Degrelle SA, Gómez E, et al. (2012). Efficient derivation of bovine embryonic stem cells needs more than active core pluripotency factors. Mol. Reprod. Dev. 79: 461-477. https://doi.org/10.1002/mrd.22051
- McDaniel BT, Miller RH and Courley EL. (1965). DHIA factors for projecting incomplete records to 305 days. Dairy Herd Improvement Letter 41, No. 6. U.S. Dept. Agr. ARS 44-164, 21.
- Norman HD, Miller PD, McDaniel BT, Dickinson FN et al. (1974). USDA-DHIA factors for standardizing 305 day lactation records for age and moth to calving. U.S. Dept. Agr. ARS-NE-40.
- NRC (2001). Nutrient requirements of dairy cattle. 7th rev. ed. Natl. Acad. Press Washington, DC.
- Nussey SS, Prysor-Jones RA, Taylor A, Ang VTY, et al. (1987). Arginine vasopressin and oxytocin in the bovine adrenal gland. J. Endocrinol. 115: 141-149.https://doi.org/10.1677/joe.0.1150141
- Olson TA, Lucena C, Chase CC and Hammond AC (2003). Evidence of a major gene influencing hair length and heat tolerance in Bos taurus cattle. J. Anim. Sci. 81: 80-90. https://doi.org/10.2527/2003.81180x
- Posner SF, Vaslet CA, Jurofcik M, Lee A, et al. (2004). Stepwise posttranslational processing of progrowth hormone-releasing hormone (proGHRH). polypeptide by furin and PC1. Endocrine 23: 199-213. https://doi.org/10.1385/endo:23:2-3:199
- Raven LA, Cocks BG, Pryce JE, Cottrell JJ, et al. (2013). Genes of the RNASE5 pathway contain SNP associated with milk production traits in dairy cattle. Genet. Sel. Evol. 45: 25. https://doi.org/10.1186/1297-9686-45-25
- Rivera-Acuña F, Prado-Martínez E, Luna-Nevárez P, Méndez-Castillo MG, et al. (2015). Induction of lactation in Holstein cows using progesterone injections or progesterone vaginal inserts. Iranian J. Appl. Anim. Sci. 5: 13-20.
- Shaji S, Niyas PAA, Chaidanya K, Sejian V, et al. (2015) Ameliorative strategies to sustain livestock production during heat stress. J. Vet. Sci. Med. Diagn. 4: 3.https://doi.org/10.4172/2325-9590.1000161
- West JW (2003). Effects of heat-stress on production in dairy cattle. J. Dairy Sci. 86: 2131-2144. https://doi.org/10.3168/jds.s0022-0302(03)73803-x
- Wiggans GR. 1985. National buck evaluations. National Cooperative Dairy Herd Improvement Program Handbook, Fact Sheet L-2. Extension Service, USDA, Washington, DC.
- Zhang J, Zan L, Fang P, Zhang F, et al. (2008). Genetic variation of PRLR gene and association with milk performance traits in dairy cattle. Can. J. Anim. Sci. 88: 33-39. https://doi.org/10.4141/cjas07052
Keywords:
Download:
Full PDF- Share This